How do we know what we don’t know? ¿Cómo sabemos lo que no sabemos?

This talk is about visualisation to help in clustering high-dimensional data
The greatest value of a data plot is when it forces us to notice what we never expected to see. ~Adapted from a Tukey quote.
Outline
- Become familiar with tour for viewing high dimensions
- Spin-and-brush
- More details of tours
- Related methods
- Clustering and tours
- Model-based clustering
- Summarising clusters
- Comparing methods
- Dimension reduction
- What we’d like to do: future research topics
Avoid cherry picking, look at all

Image: Sketchplanations
High-dimensional visualisation
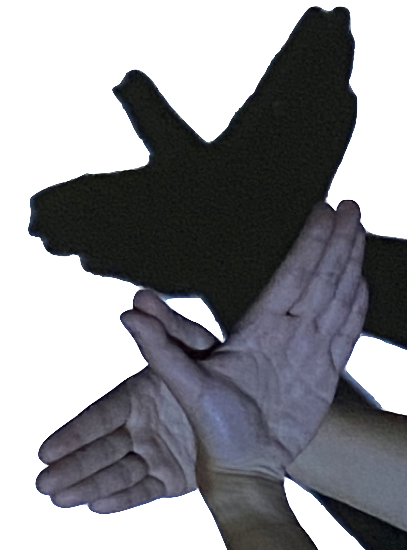

Tours of high-dimensional data are like examining the shadows (projections)
(and slices/sections to see through a shadow)
Notation
Data
Notation
Projection
Notation
Projected data
High-dimensional visualisation
Data is 2D:
Projection is 1D:
Notice that the values of change between (-1, 1). All possible values being shown during the tour.
watching the 1D shadows we can see:
- unimodality
- bimodality, there are two clusters.
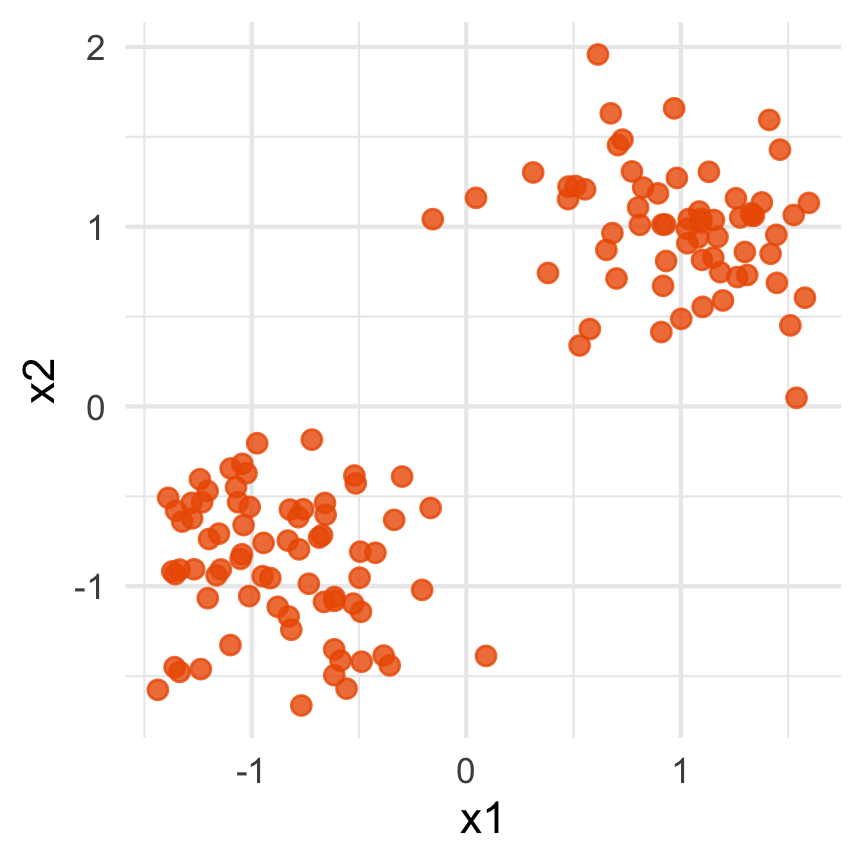
What does the 2D data look like? Can you sketch it?
High-dimensional visualisation
⟵
The 2D data
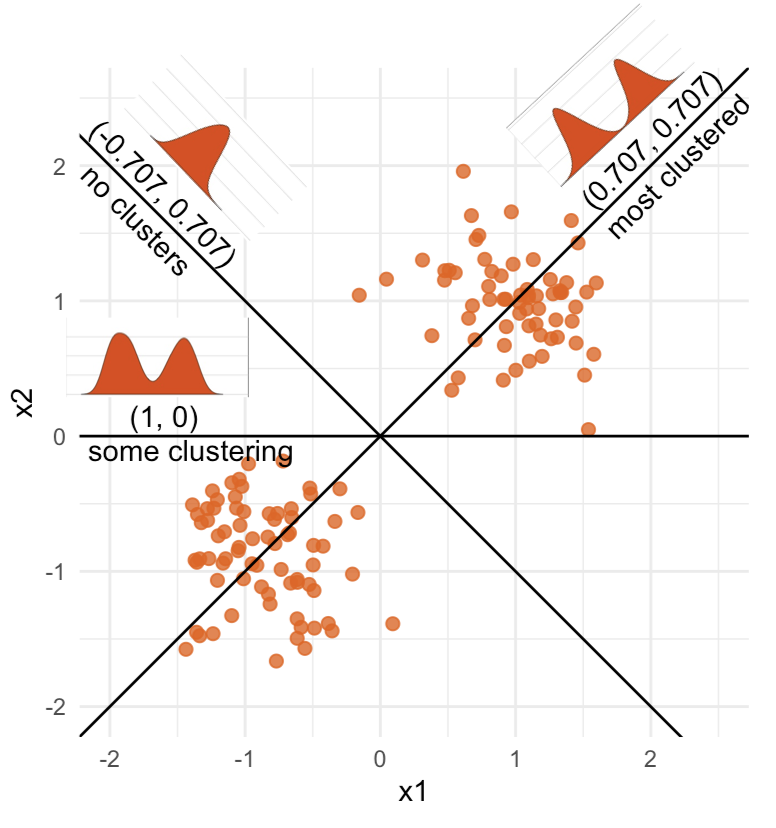
High-dimensional visualisation

Data is 3D:
Projection is 2D:
Notice that the values of change between (-1, 1). All possible values being shown during the tour.
See:
- circular shapes
- some transparency, reveals middle
- hole in in some projections
- no clustering
High-dimensional visualisation
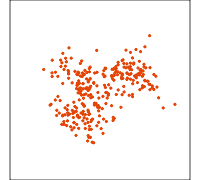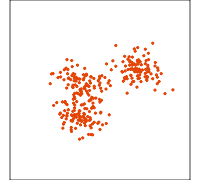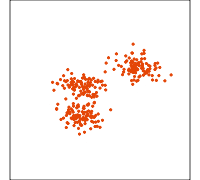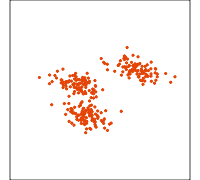
Data is 4D:
Projection is 2D:
How many clusters do you see?
- three, right?
- one separated, and two very close,
- and they each have an elliptical shape.
- do you also see an outlier or two?
Well done!
You can now tell everyone that you can SEE in 4D!
Method 1: Spin-and-brush
If you want to discover and mark the clusters you see, you can use the detourr package to spin and brush points. Here’s a live demo. Hopefully this works.
Tour architecture
- Data: -D
- Projection dimension: choose
- Rendering method: histogram, density plot, scatterplot, …
Algorithm:
- Path taken through high-dimensions: random, guided, local, little, manual
- Interpolation method: geodesic (plane to plane), Givens (basis to basis)
Software:
Types of tours
Grand tour
Slice display
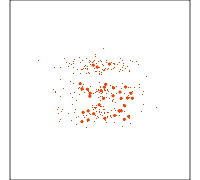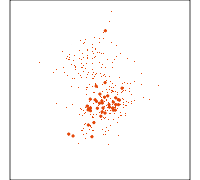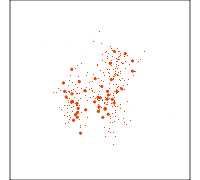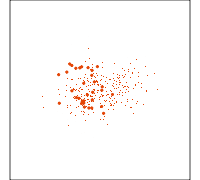
Guided tour
Local tour
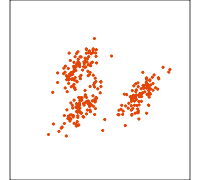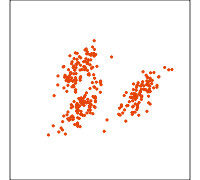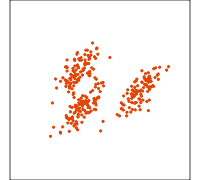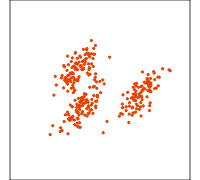
Related methods
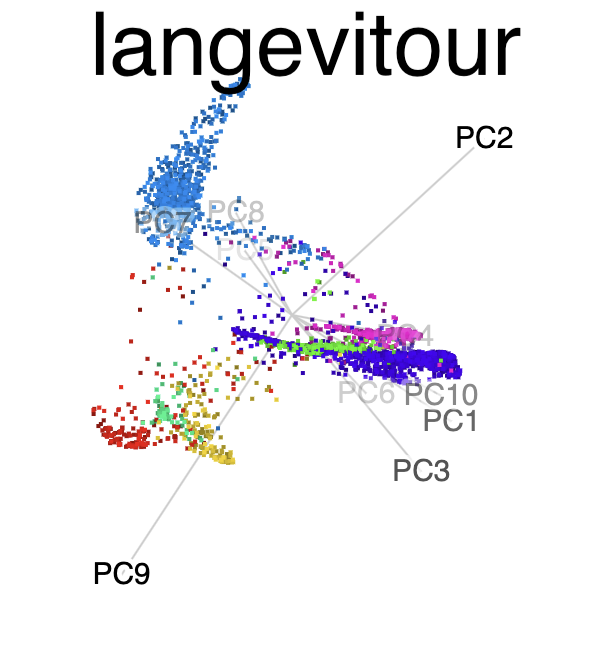
PCA
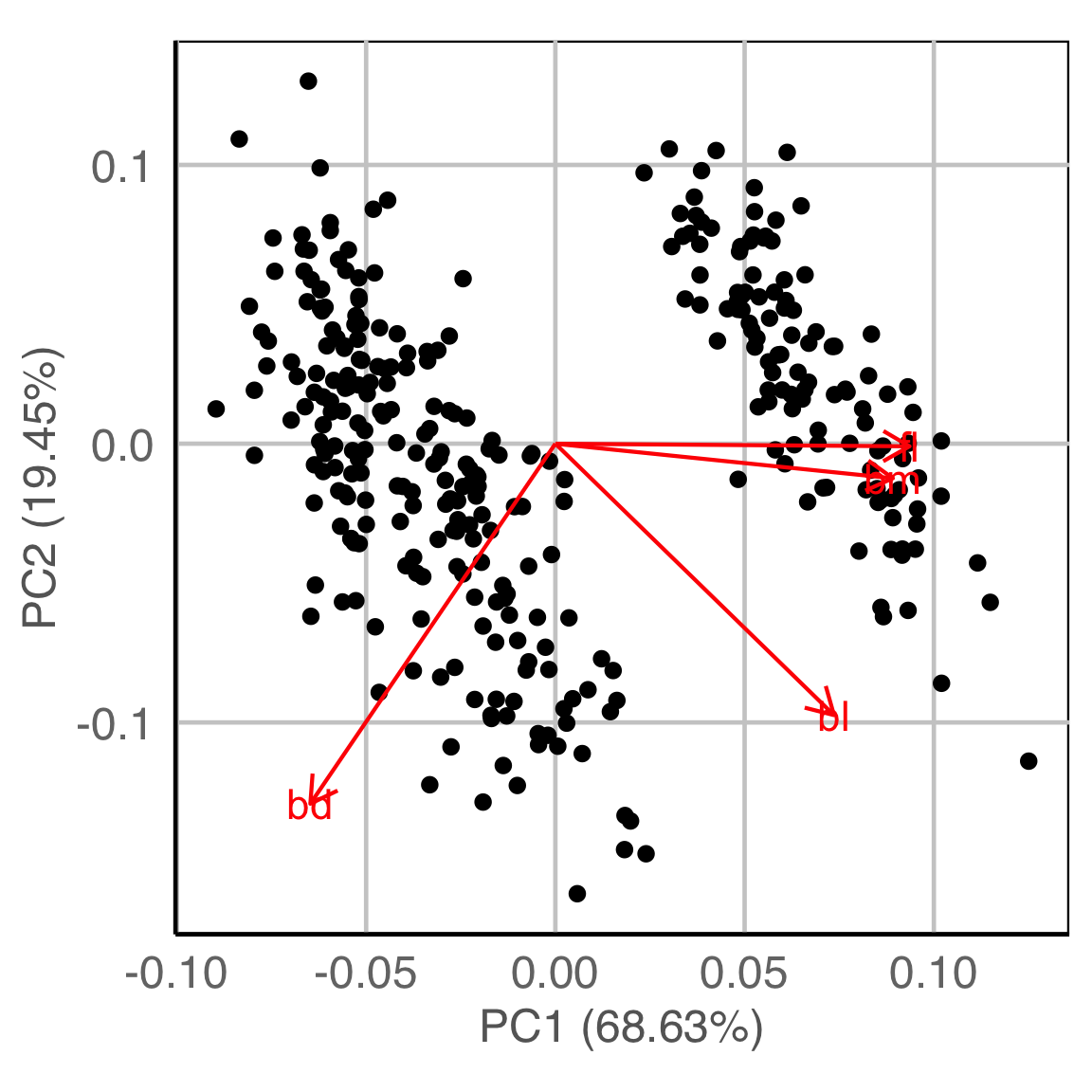
NLDR: tSNE
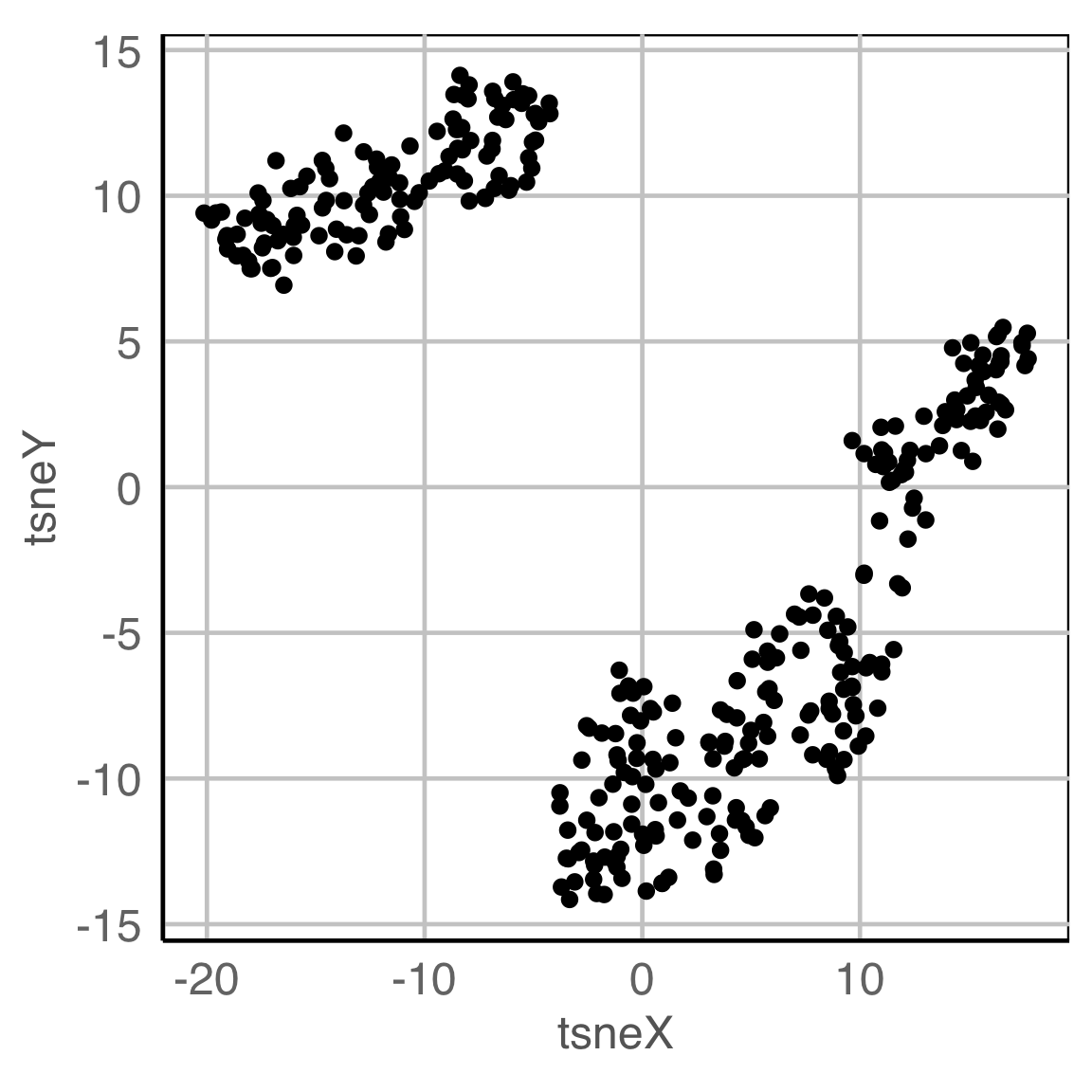
Clustering & tours
Model-based - 2D (1/3)
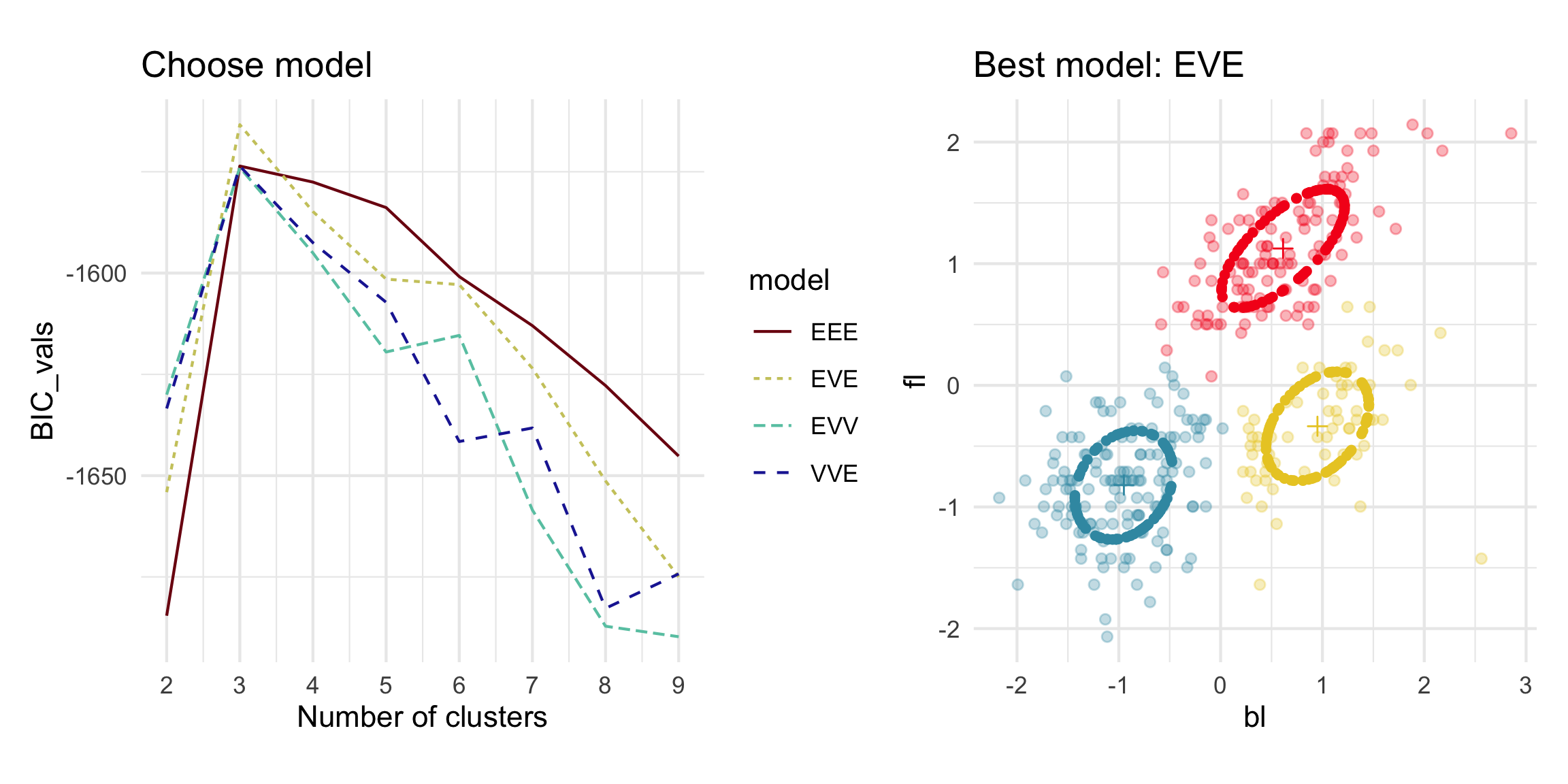
Model-based - 4D (2/3)
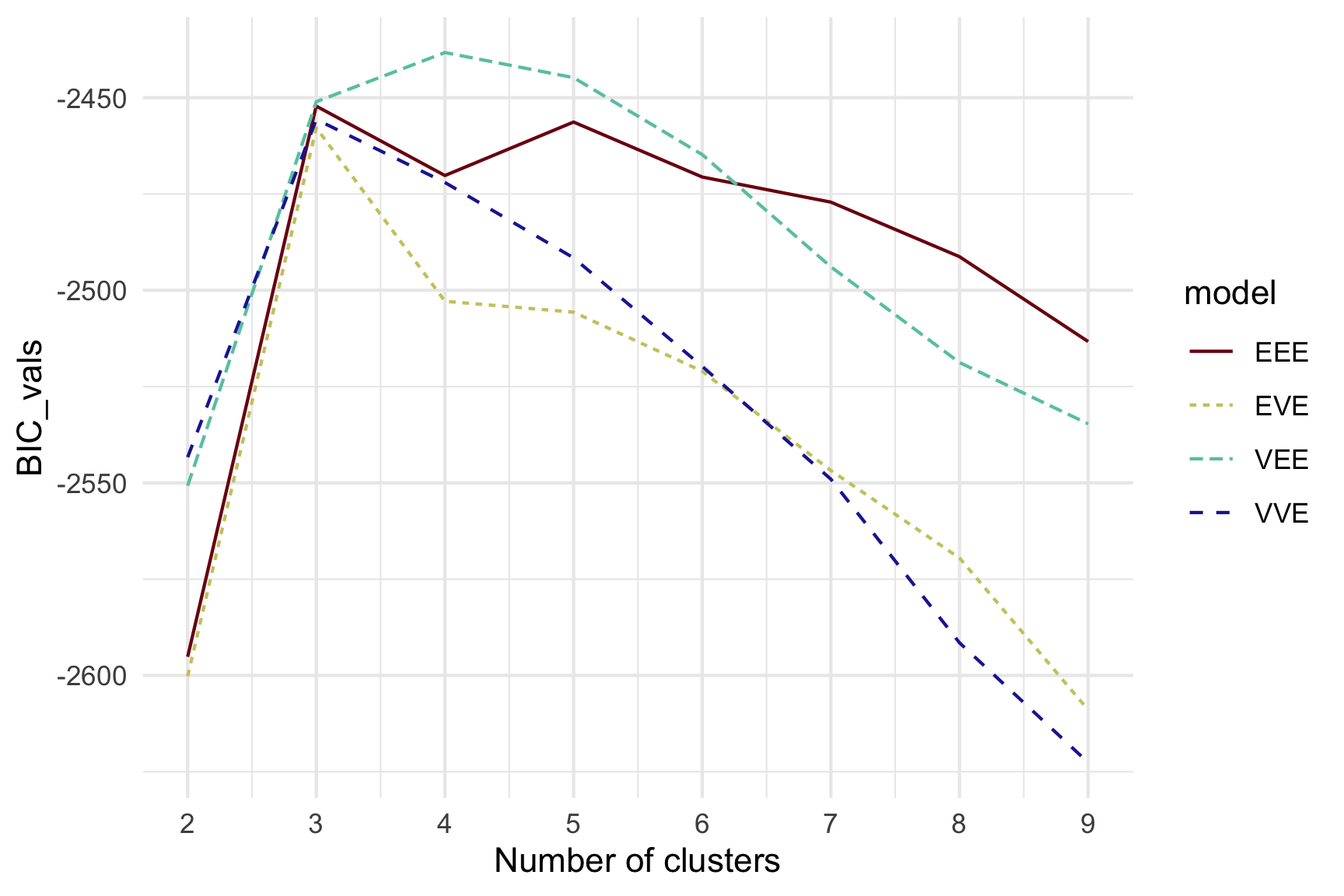
Model-based (3/3) ~~Which fits the data better?
Best model: four-cluster VEE
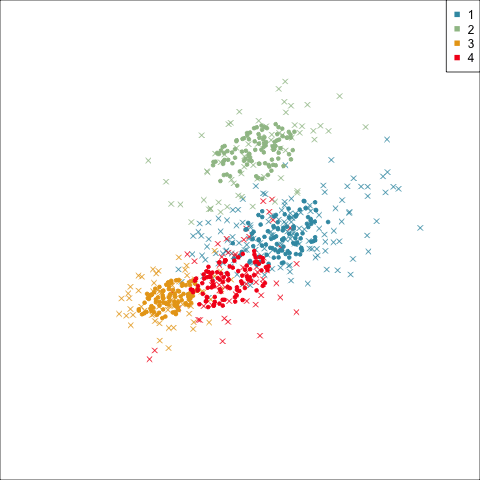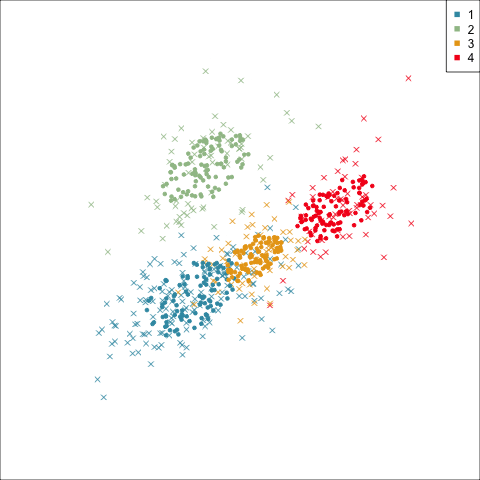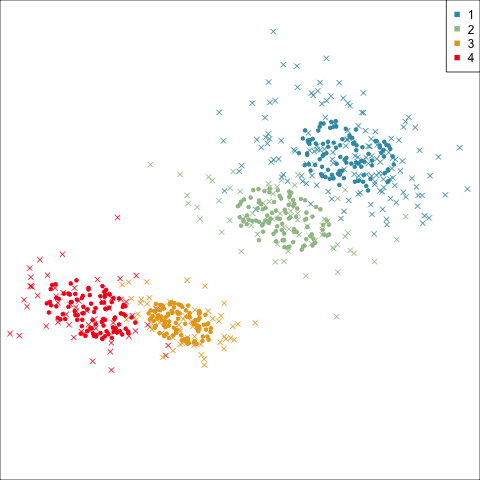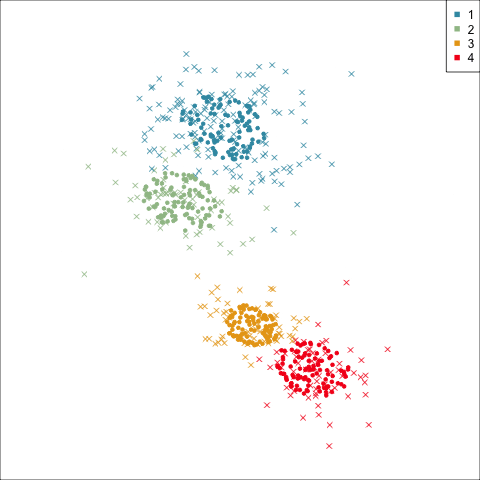
Three-cluster EEE
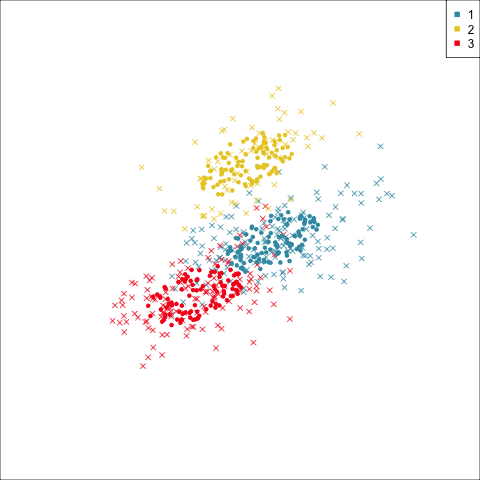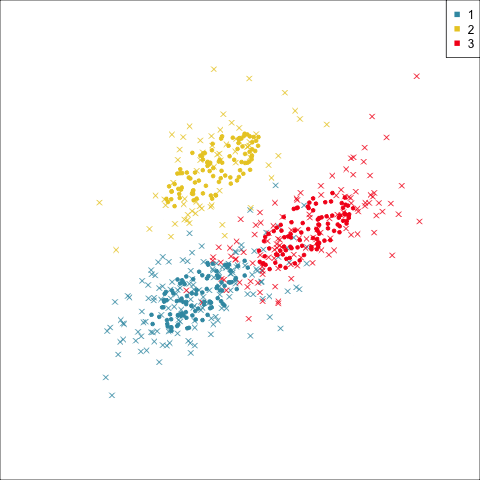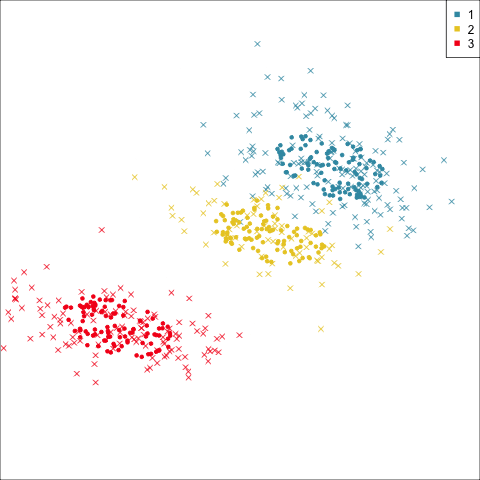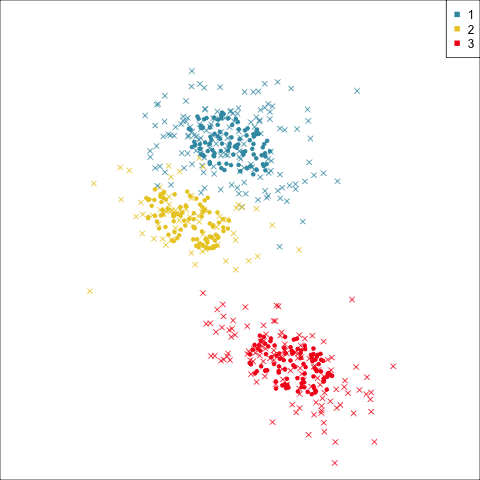
Summarising clusters
Convex hulls are often used to summarise clusters in 2D. It is possible to view these in high-d, too.
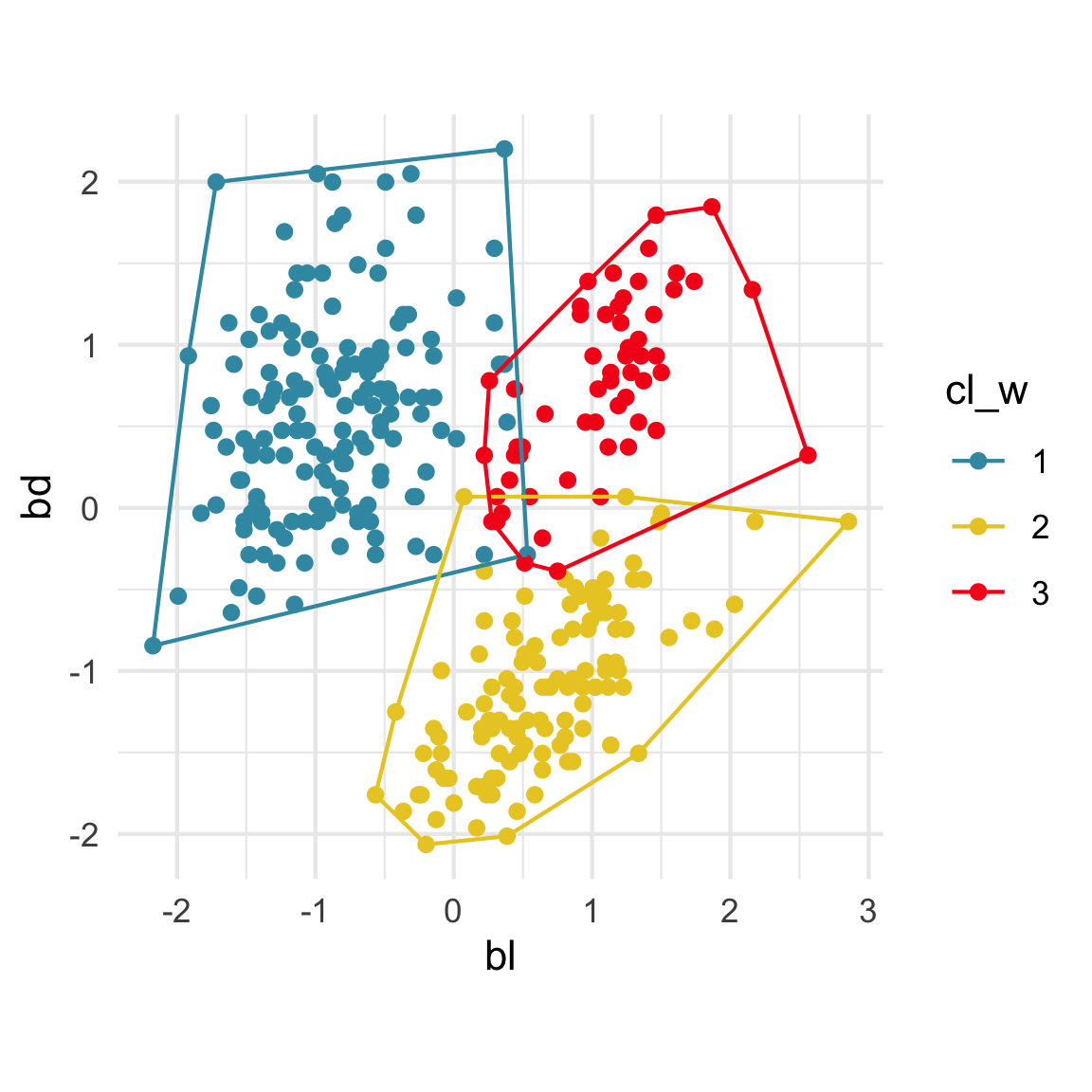
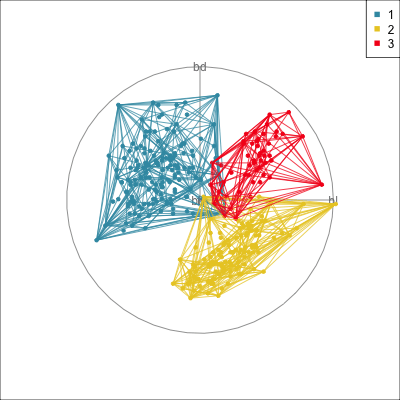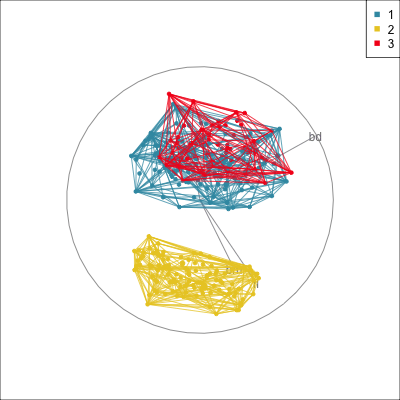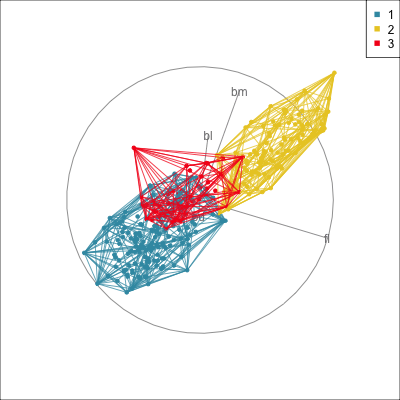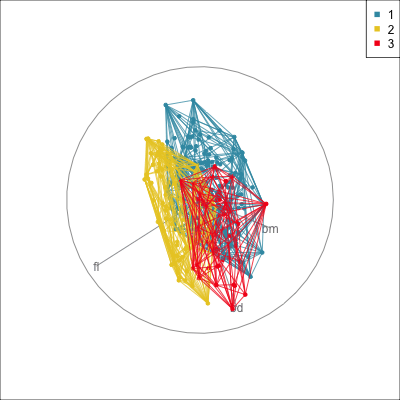
Comparing methods
| cl_w | cl_mc | ||
|---|---|---|---|
| 1 | 2 | 3 | |
| 1 | 149 | 8 | 0 |
| 2 | 0 | 0 | 119 |
| 3 | 0 | 57 | 0 |
library(crosstalk)
library(plotly)
library(viridis)
p_cl_shared <- SharedData$new(penguins_cl)
detour_plot <- detour(p_cl_shared, tour_aes(
projection = bl:bm,
colour = cl_w)) |>
tour_path(grand_tour(2),
max_bases=50, fps = 60) |>
show_scatter(alpha = 0.7, axes = FALSE,
width = "100%", height = "450px")
conf_mat <- plot_ly(p_cl_shared,
x = ~cl_mc_j,
y = ~cl_w_j,
color = ~cl_w,
colors = viridis_pal(option = "D")(3),
height = 450) |>
highlight(on = "plotly_selected",
off = "plotly_doubleclick") %>%
add_trace(type = "scatter",
mode = "markers")
bscols(
detour_plot, conf_mat,
widths = c(5, 6)
) Dimension reduction
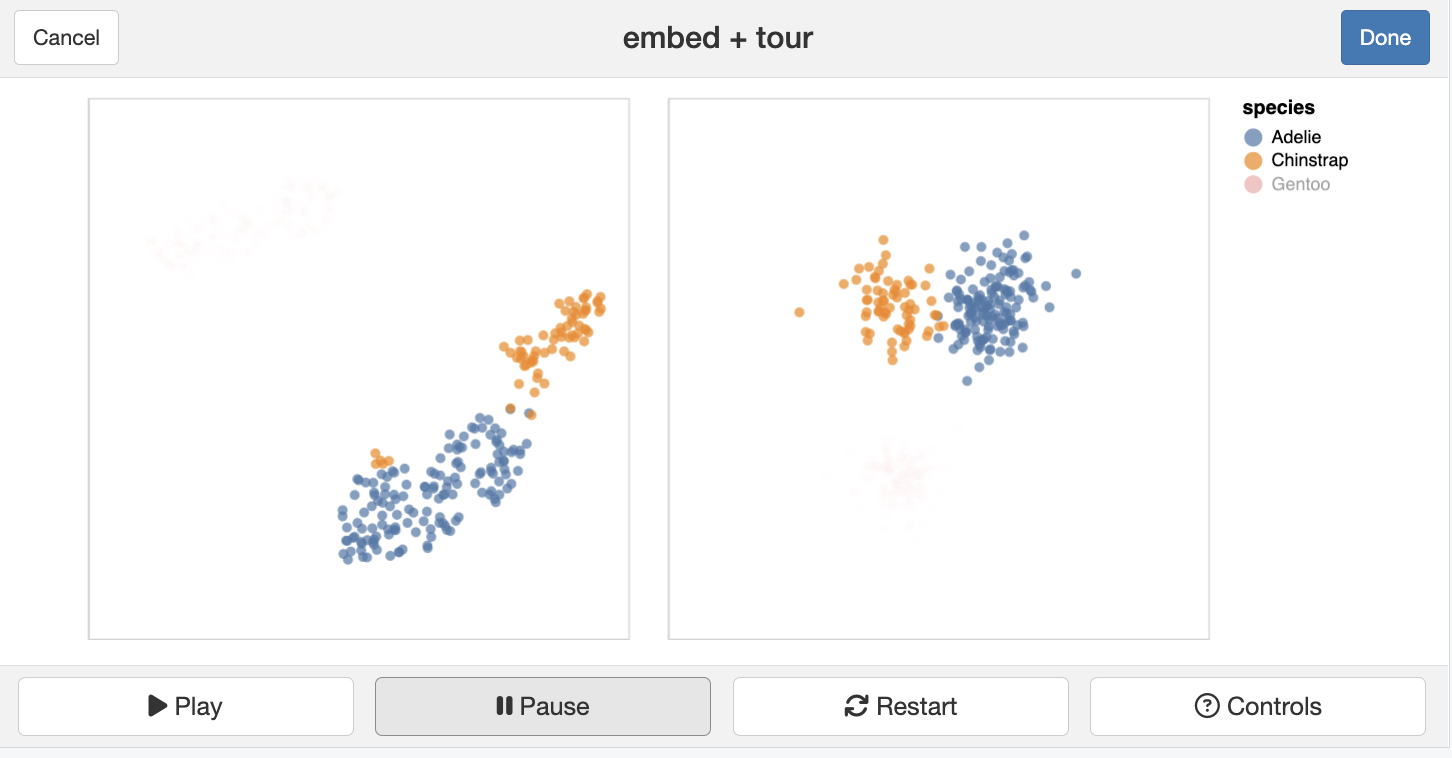
What we can’t do, that we’d like to
The tourr package provides the algorithm to generate the tour paths, and also create new tours, different displays. However, the interactivity is poor, which is a big limitation.
- Stopping, pausing, going back
- Zooming in, focus on subsets
- Linking between multiple displays
detourr is an elegant solution, which could be developed further.
- Better integration with model objects
- Specialist design for different models
- Integrating other guidance, explainability metrics
Vis is fun!
References and acknowledgements
- Cook and Laa (2023) Interactively exploring high-dimensional data and models in R
- Slides made in Quarto.
- Get a copy of slides at https://github.com/dicook/LatinR

This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.