New tools for visualising high-dimensional data using linear projections
Econometrics and Business Statistics
Monash University
You can’t see beyond 3D!
You can’t see beyond 3D!
We are going to see that we can gain intuition for structure in high dimensions through visualisation
The greatest value of a data plot is when it forces us to notice what we never expected to see. ~Adapted from a Tukey quote.
It doesn’t mean that it’s easy. It doesn’t mean that visualisation is used alone. It means that (high-dimensional) visualisation is an important part of your toolbox, especially to allow discovery of what we don’t know.
Outline
- Using a tour to see into high dimensions
- Algorithms in the tourr package
- New developments in recent years
- Examples of usage
- Future research directions
- Other talks at this conference
High-dimensional visualisation

Tours of high-dimensional data are like examining the shadows (projections)
(and slices/sections to see through a shadow)
High-dimensions in statistics
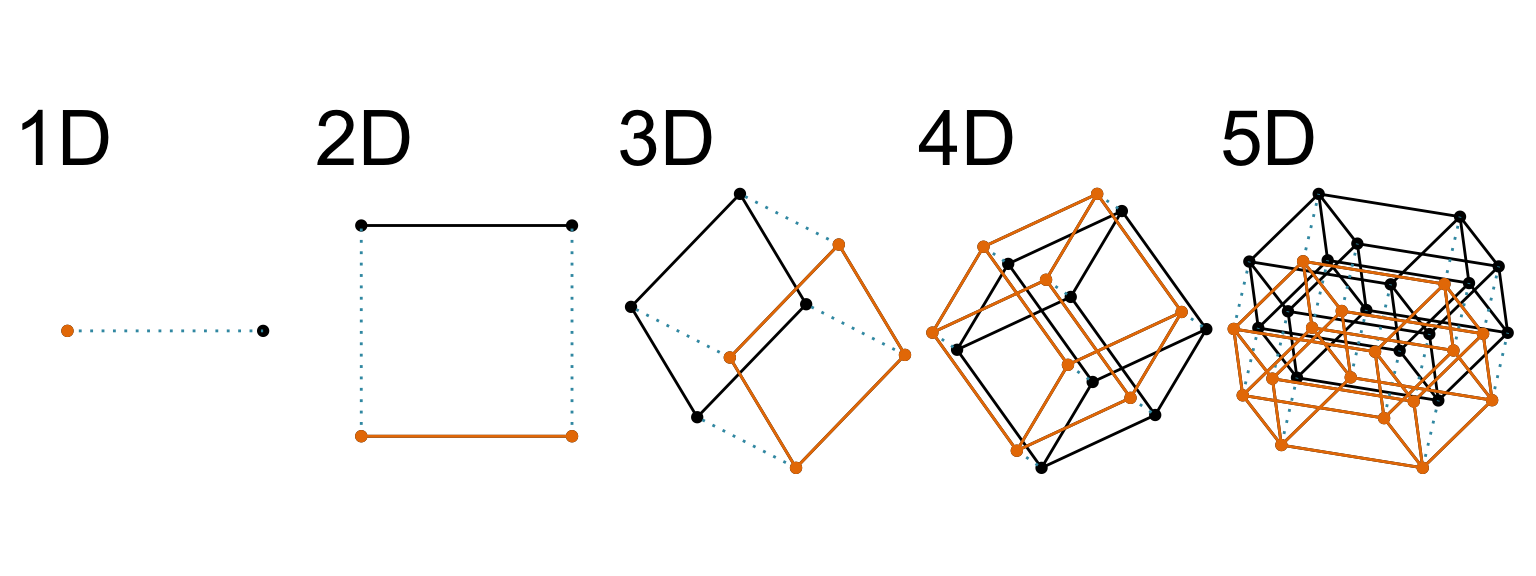
Increasing dimension adds an additional orthogonal axis.
If you want more high-dimensional shapes there is an R package, geozoo, which will generate cubes, spheres, simplices, mobius strips, torii, boy surface, klein bottles, cones, various polytopes, …
And read or watch Flatland: A Romance of Many Dimensions (1884) Edwin Abbott.
Explanation
Data
Explanation
Projection
Explanation
Projected data
High-dimensional visualisation
Data is 2D:
Projection is 1D:
Notice that the values of change between (-1, 1). All possible values being shown during the tour.
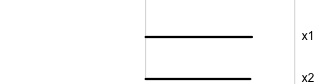
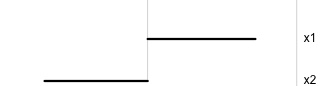
watching the 1D shadows we can see:
- unimodality
- bimodality, there are two clusters.
What does the 2D data look like? Can you sketch it?
High-dimensional visualisation
⟵
The 2D data
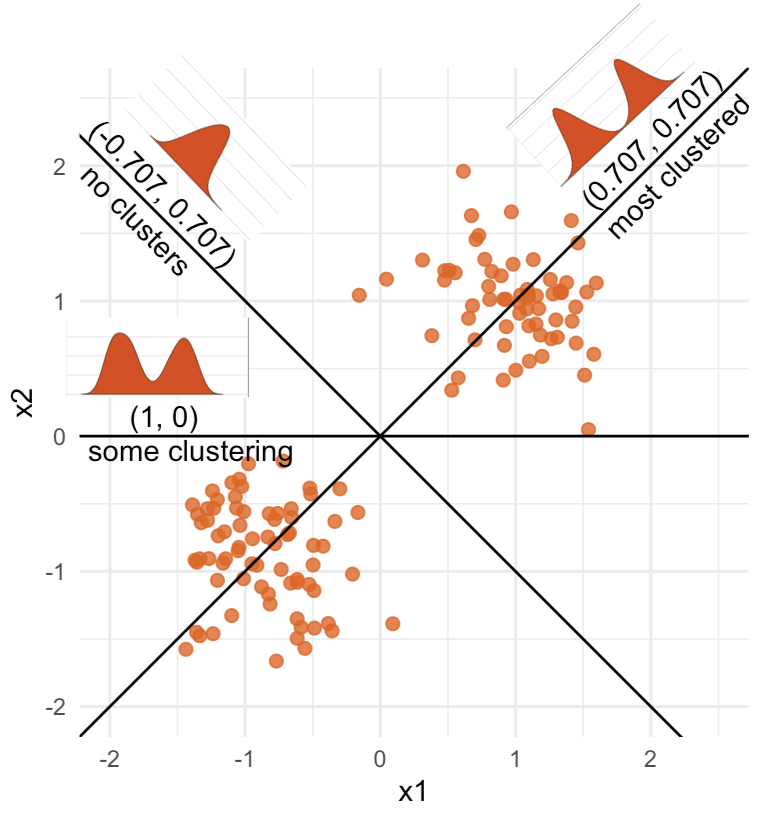
High-dimensional visualisation

Data is 3D:
Projection is 2D:
Notice that the values of change between (-1, 1). All possible values being shown during the tour.
See:
- circular shapes
- some transparency, reveals middle
- hole in in some projections
- no clustering
High-dimensional visualisation
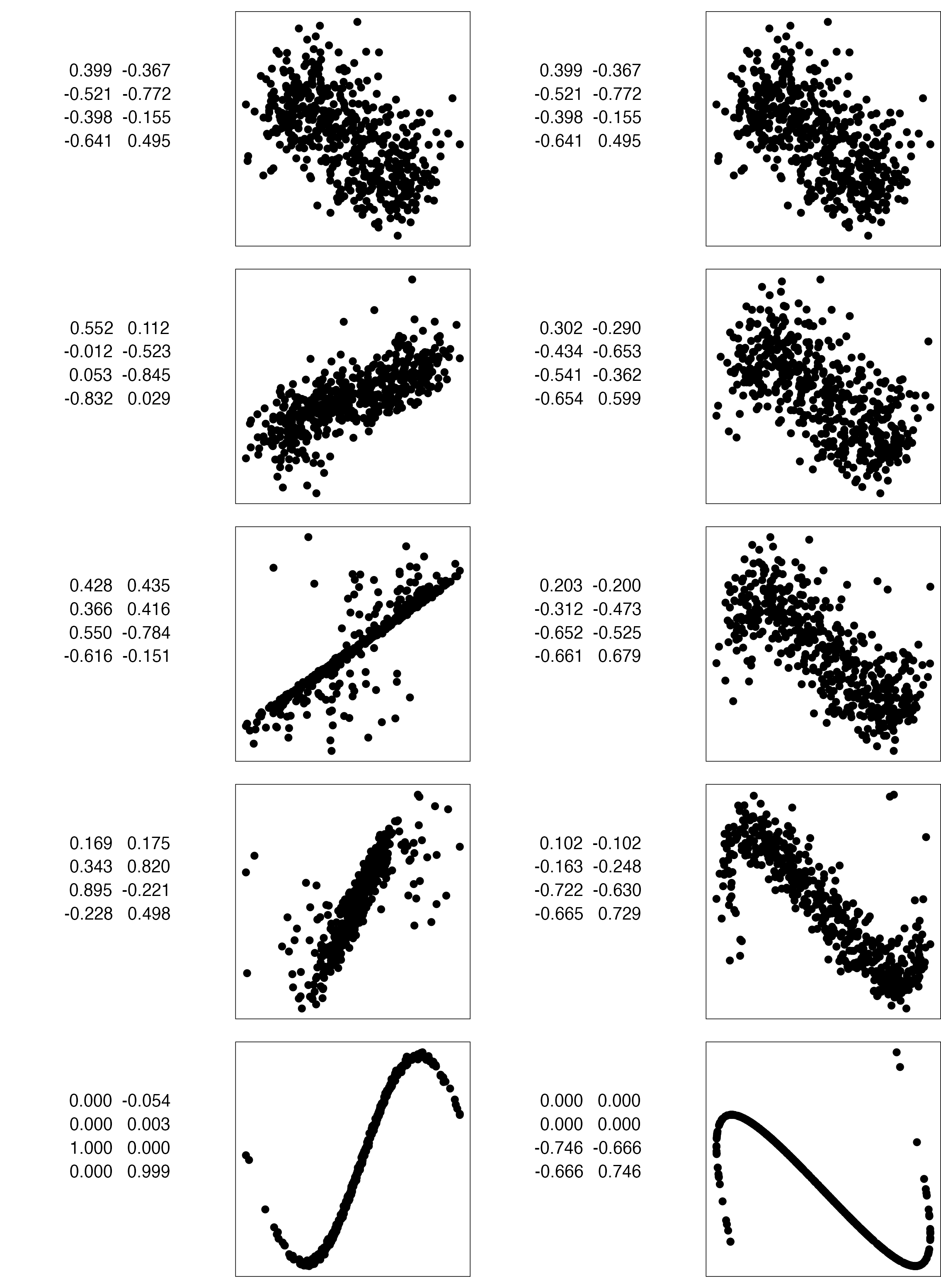
Data is 4D:
Projection is 2D:
How many clusters do you see?
- three, right?
- one separated, and two very close,
- and they each have an elliptical shape.
- do you also see an outlier or two?
Early tour algorithms

1D paths in 3D space

2D paths in 3D space
Early tour algorithms
Grand tour: see from all sides

Guided tour: Steer towards the most interesting features.
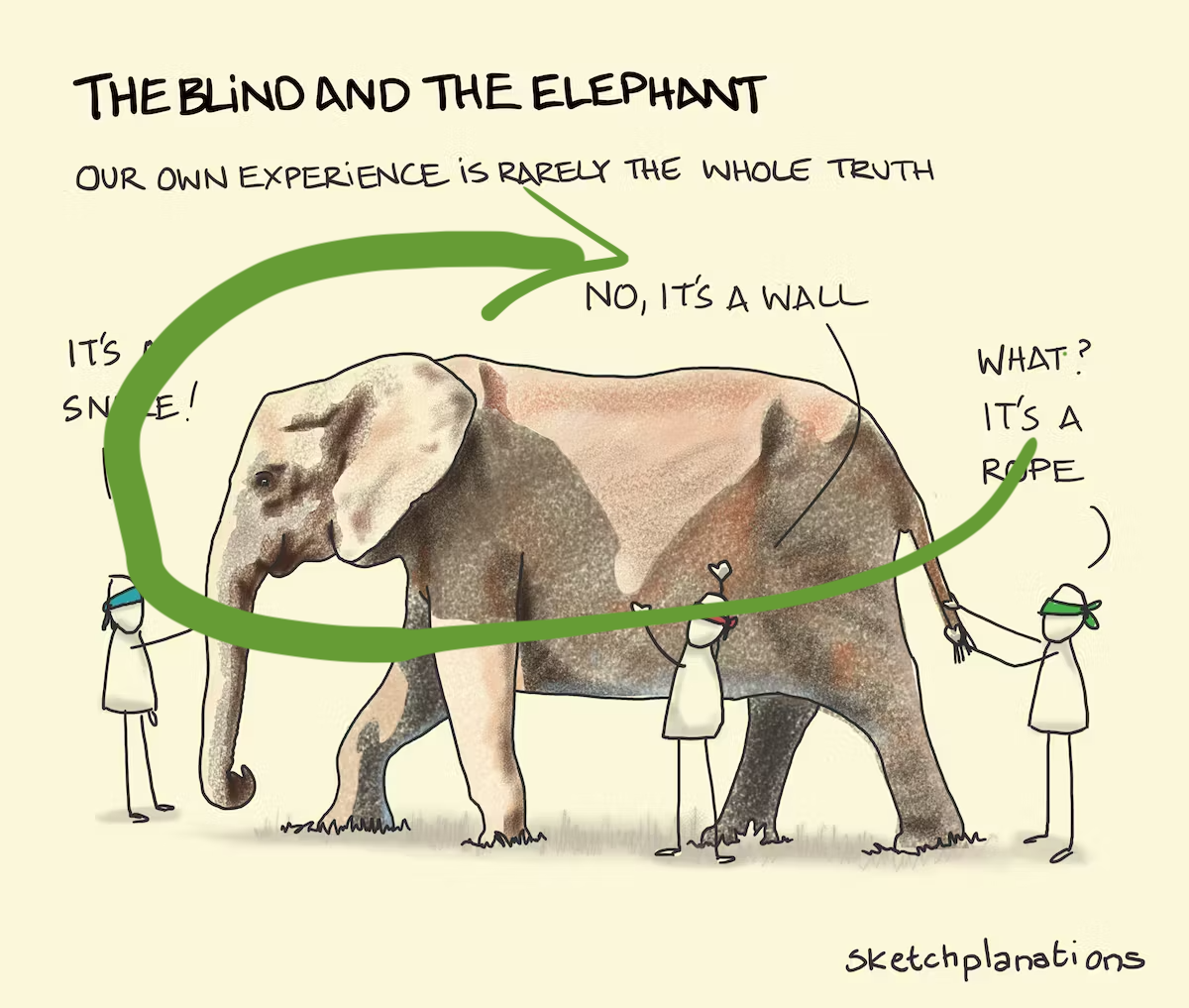
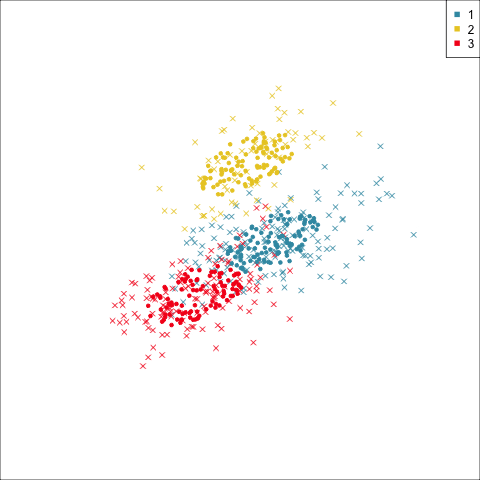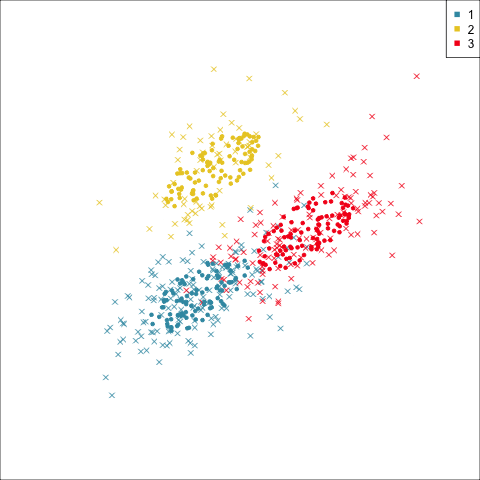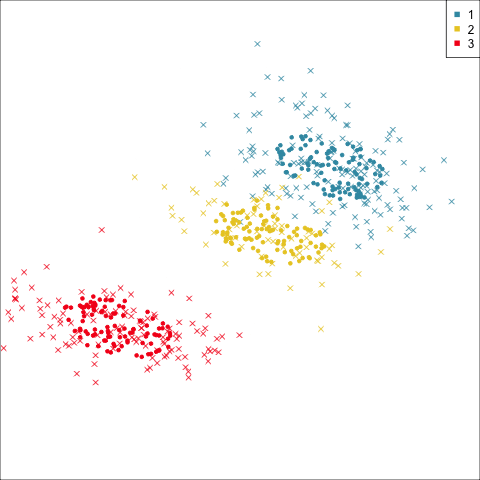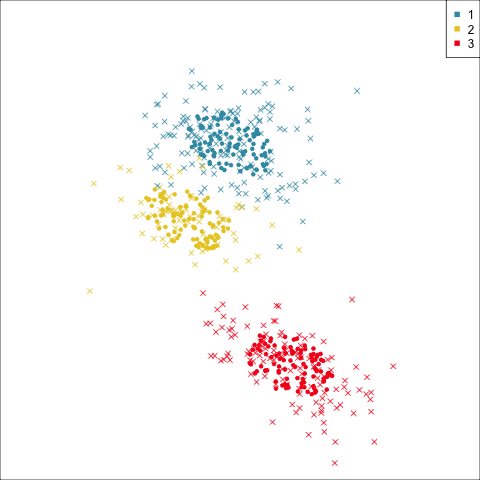
Why? (Three cluster data)
Avoid being a blind man inspecting the elephant
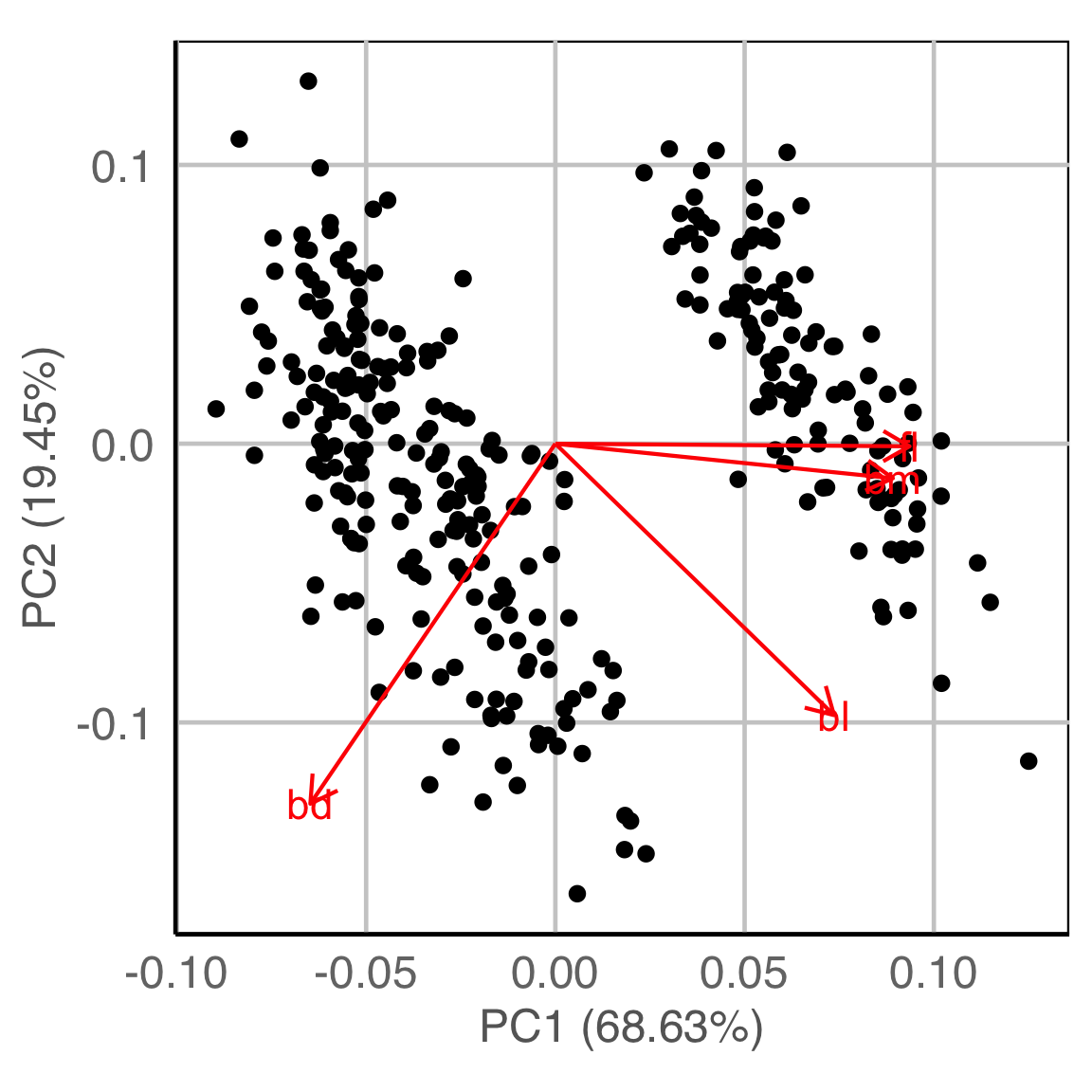
Principal component analysis
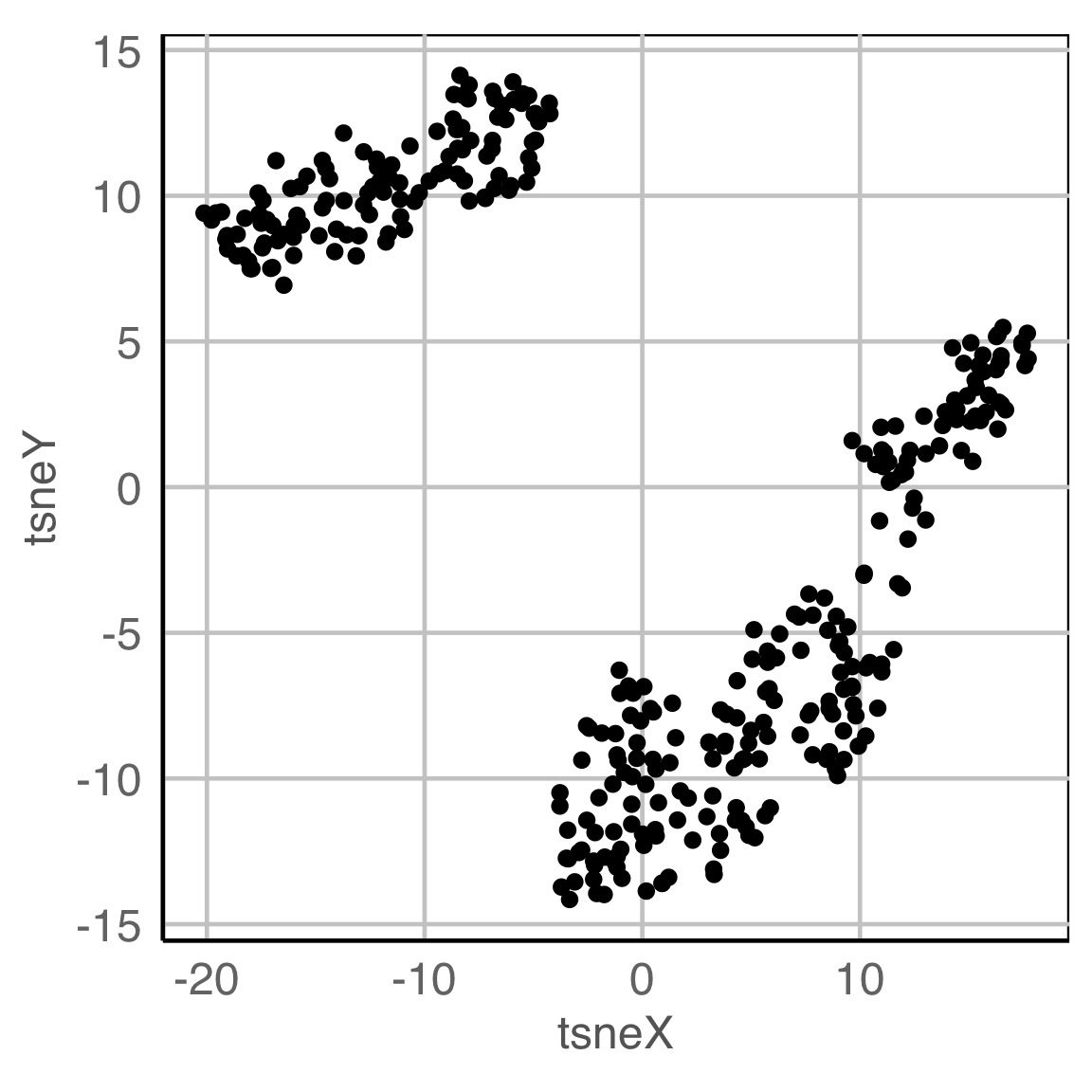
NLDR: t-Stochastic neighbourhood embedding
Algorithms in the tourr package

Movement
- choice of target planes
- grand: random
- guided: objective function
- local: nearby
- little: marginals
- manual/radial: specific variable
- interpolation between them
- geodesic: plane to plane
- Givens: frame/basis to frame/basis
Display
How should you plot your projected data?
- 1D: density, dotplot, histogram
- 2D: scatterplot, density2D, sage, pca, slice
- 3D: stereo
- kD: parallel coordinates, scatterplot matrix
- 1D+spatial: image
Recent developments
- interactivity: detourr, liminal, langevitour
- slice/section: explore shape of models
- manual/radial tour: explore sensitivity of structure to particular variables
- sage: correct for piling
- Givens interpolation: frame to frame
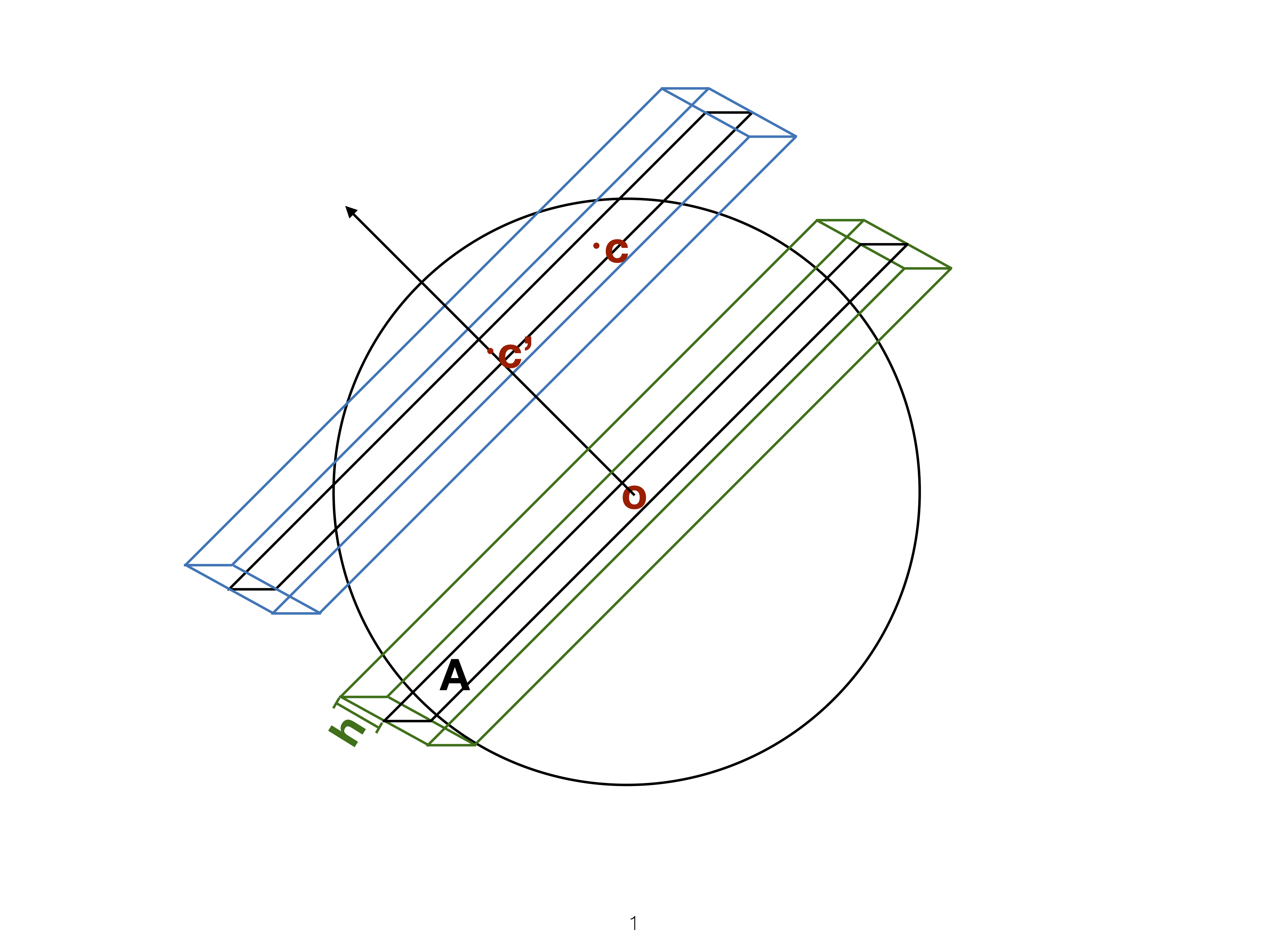
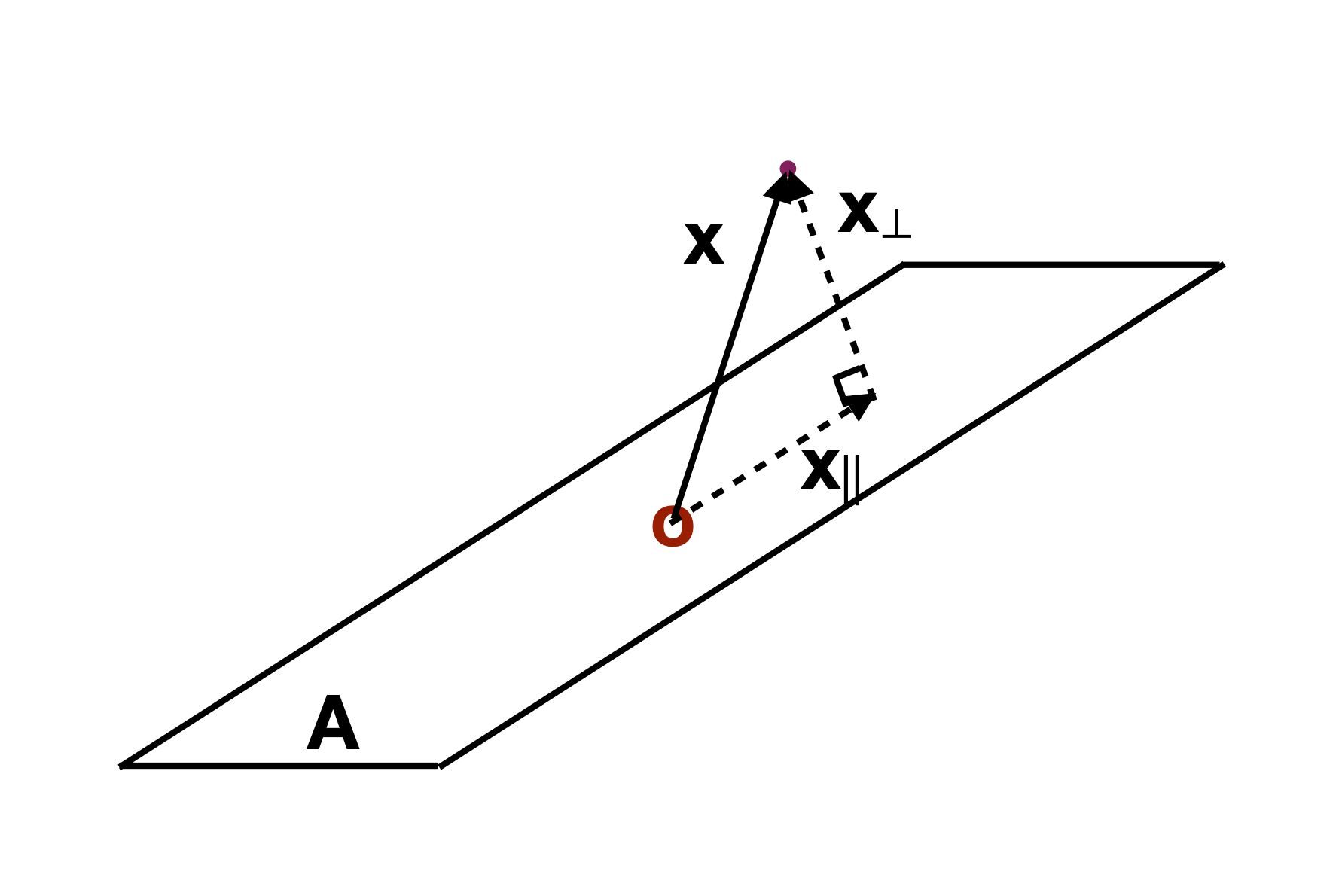
Slice
Utilise distance from the projection plane to make the slice, and shift centre of projection plane.
Sage transformation (1/2)
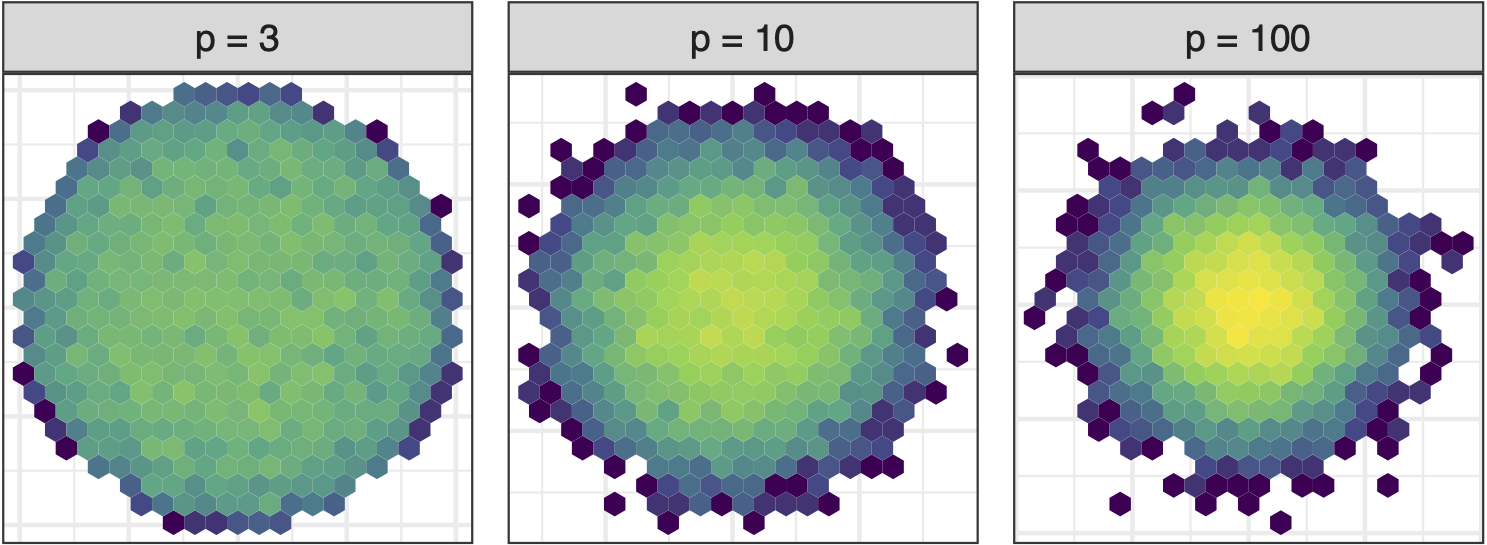
Increase variables, increase concentration, possibly obscuring important structure.
Sage transformation (2/2)
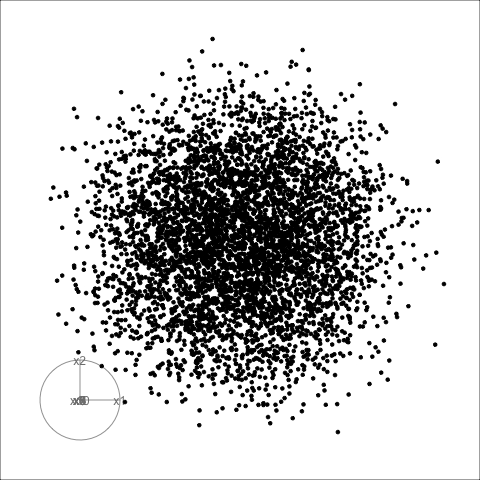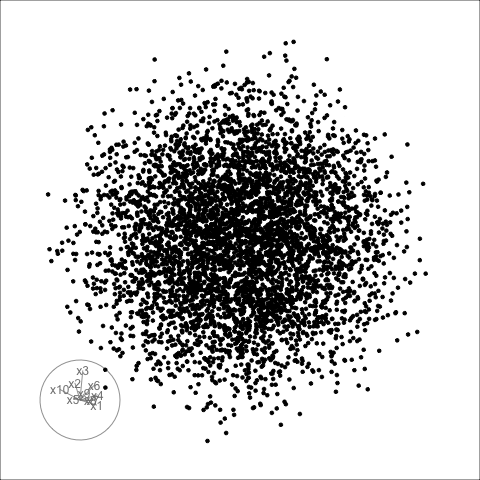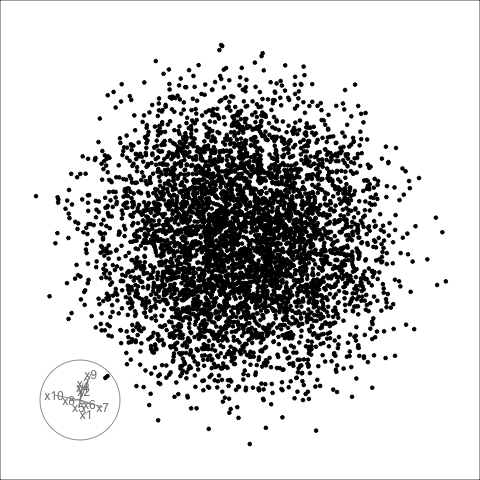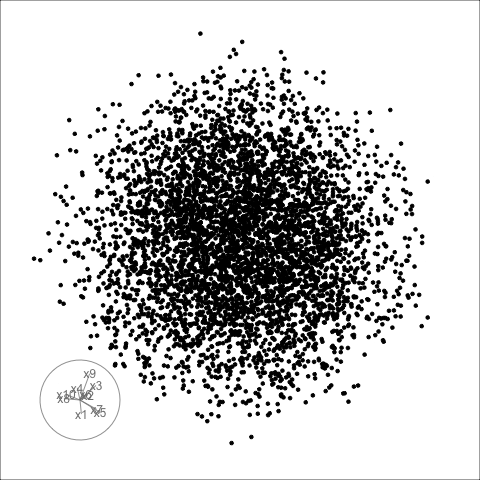

Transformation expands the centre to make a sage display.
Givens (1/2)

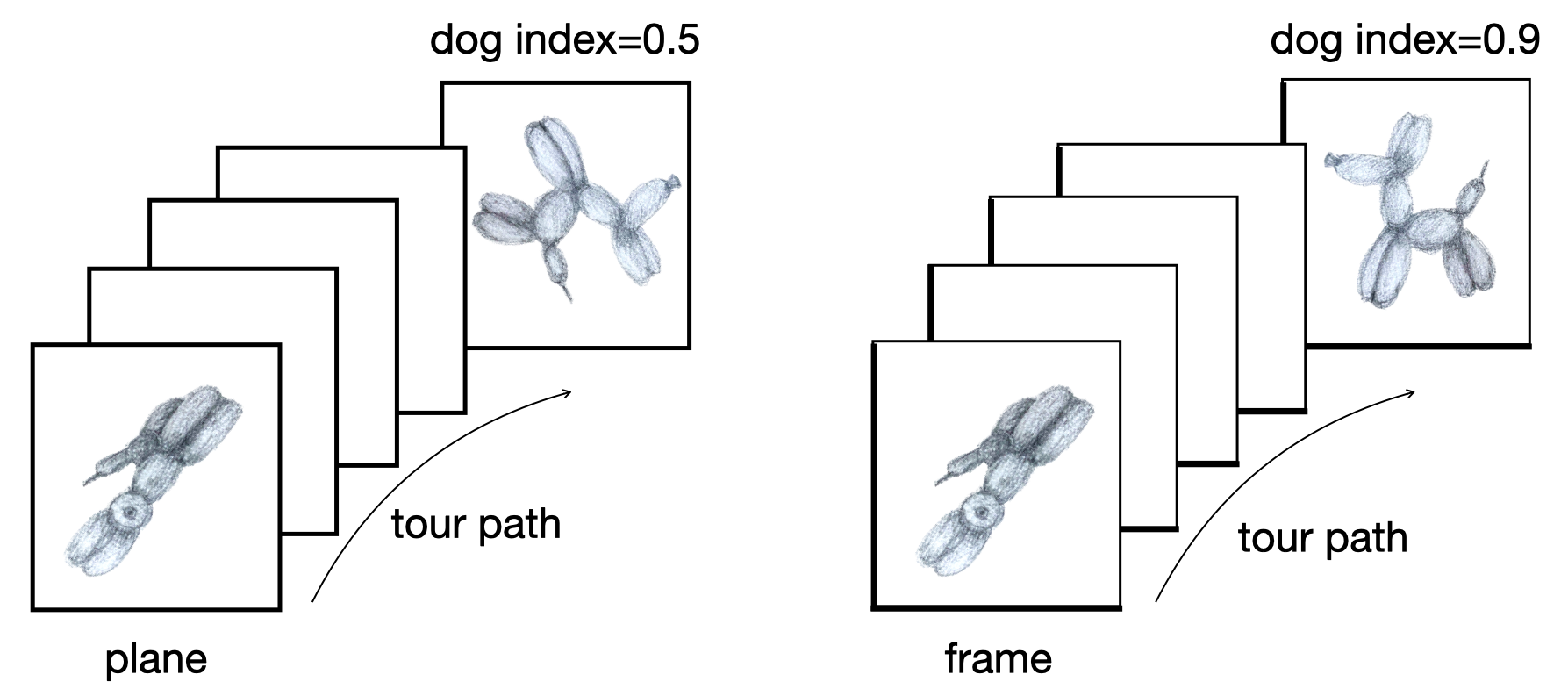
Givens (2/2)
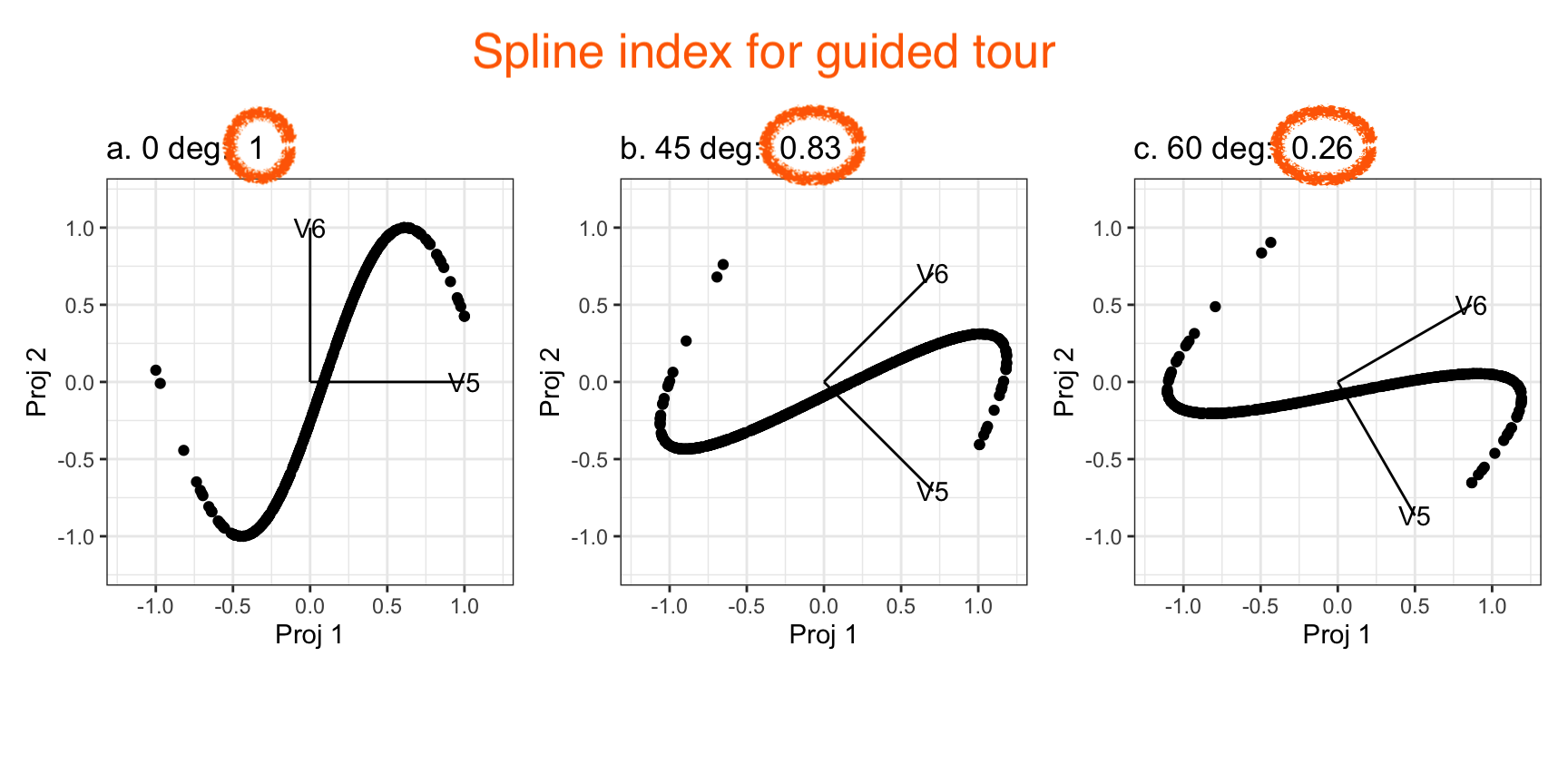
Givens interpolation ends at requested frame, but geodesic interpolation arrives at the plane, is frame-agnostic, and that is problematic for optimisation using the guided tour.
Interactivity: exploration
If you want to discover and mark the clusters you see, you can use the detourr package to spin and brush points. Here’s a live demo. Hopefully this works.
Interactivity: Compare cluster models
| cl_w | cl_mc | ||
|---|---|---|---|
| 1 | 2 | 3 | |
| 1 | 149 | 8 | 0 |
| 2 | 0 | 0 | 119 |
| 3 | 0 | 57 | 0 |
library(crosstalk)
library(plotly)
library(viridis)
p_cl_shared <- SharedData$new(penguins_cl)
detour_plot <- detour(p_cl_shared, tour_aes(
projection = bl:bm,
colour = cl_w)) |>
tour_path(grand_tour(2),
max_bases=50, fps = 60) |>
show_scatter(alpha = 0.7, axes = FALSE,
width = "100%", height = "450px")
conf_mat <- plot_ly(p_cl_shared,
x = ~cl_mc_j,
y = ~cl_w_j,
color = ~cl_w,
colors = viridis_pal(option = "D")(3),
height = 450) |>
highlight(on = "plotly_selected",
off = "plotly_doubleclick") %>%
add_trace(type = "scatter",
mode = "markers")
bscols(
detour_plot, conf_mat,
widths = c(5, 6)
) Manual/radial tour
Best projection provided by the guided tour, separating three species.
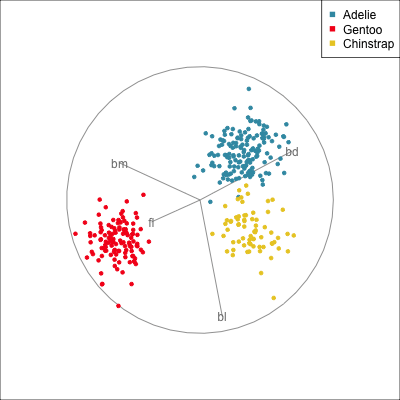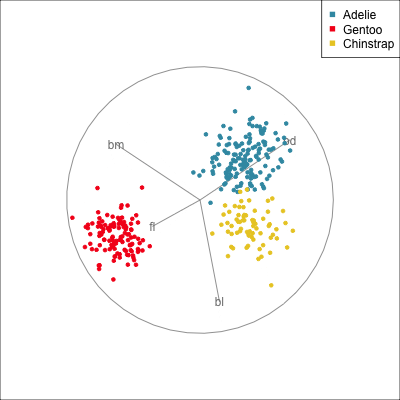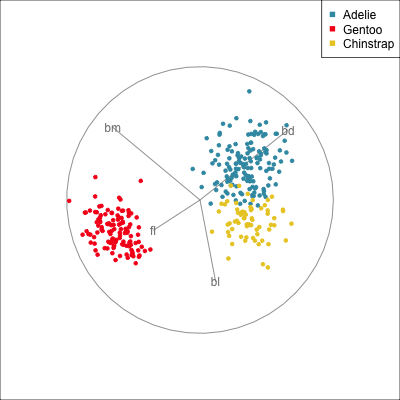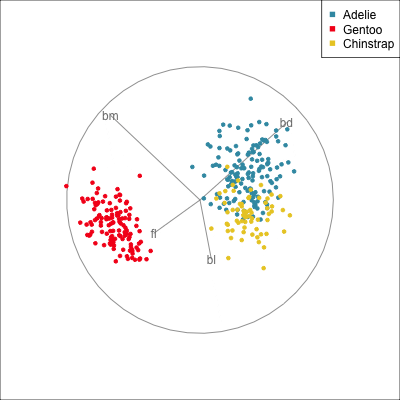
Removing flipper length
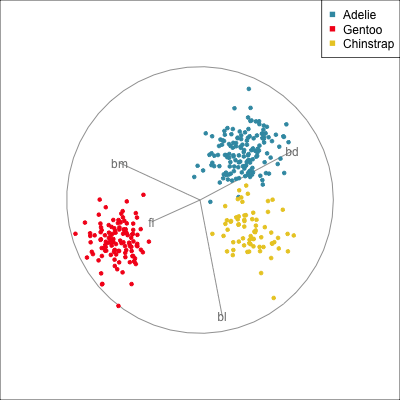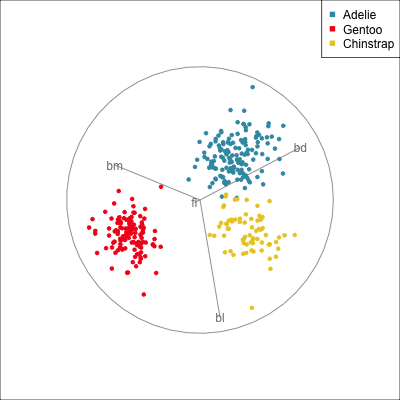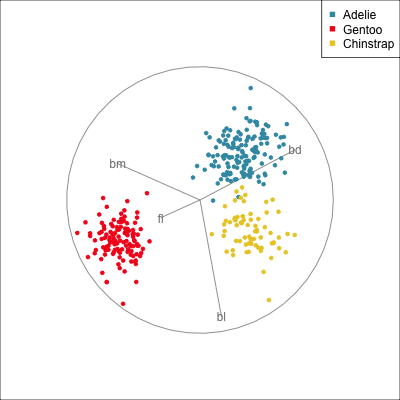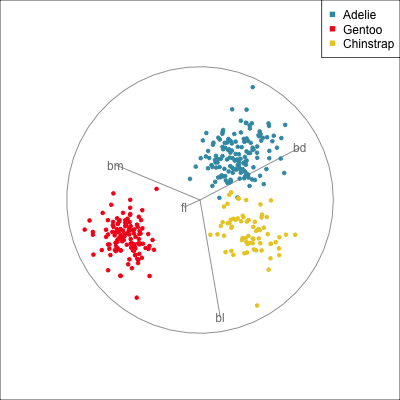
Removing bill length
Slice tour (1/2)
Projection
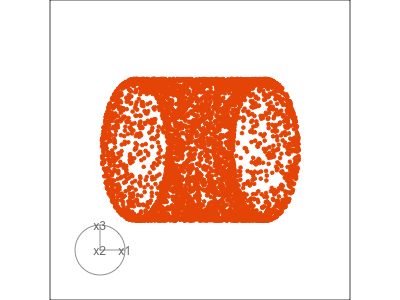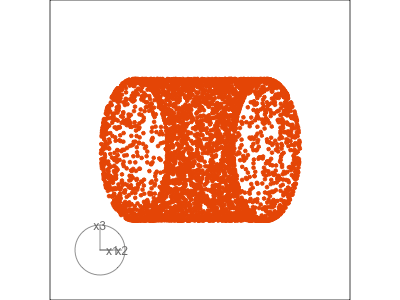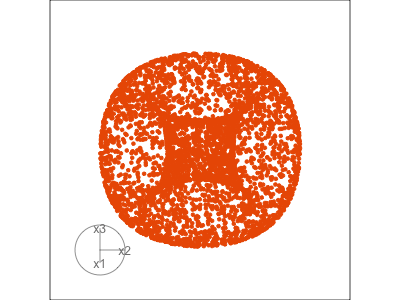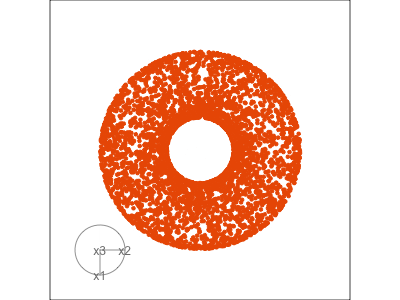
Slice

Slice tour (2/2)
This is especially useful for exploring classification models, comparing boundaries produced by different models. (The same penguins data used here.)
Linear discriminant analysis

Classification tree

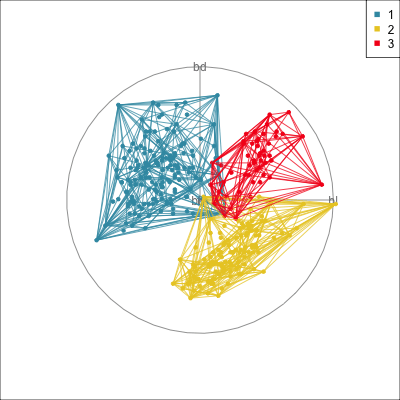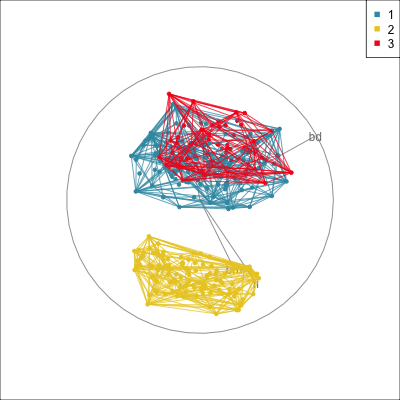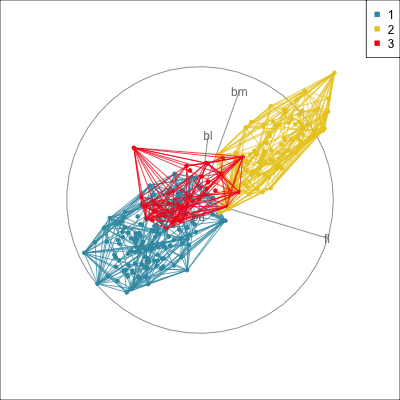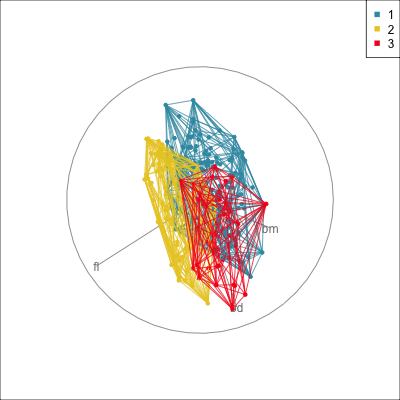
Model in the data space (1/2)
Data in the model space 1
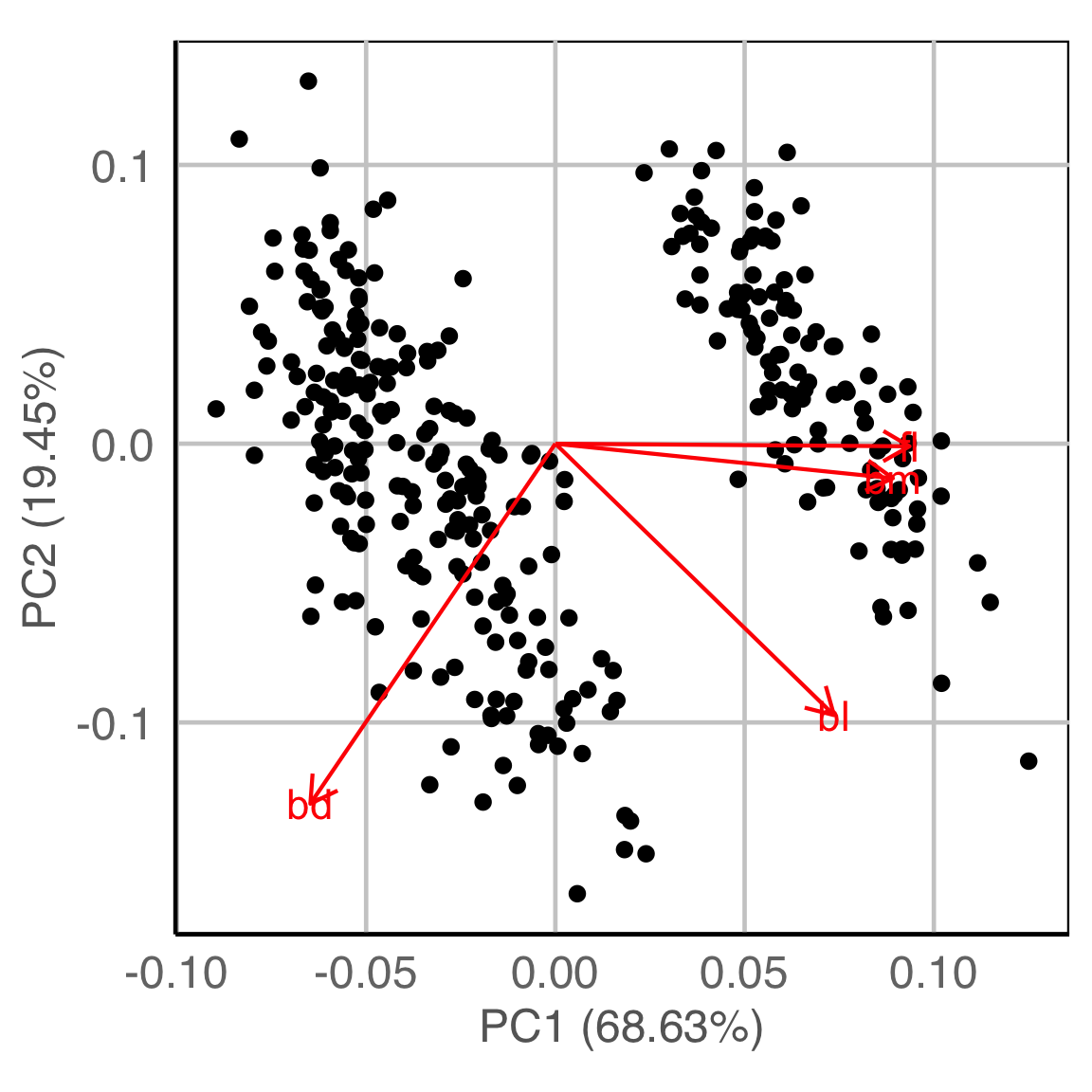
Model in the data space
Code
library(mulgar)
p_pca_m <- pca_model(p_pca, s=2.2)
p_pca_m_d <- rbind(p_pca_m$points, penguins_sub[,1:4])
animate_xy(p_pca_m_d, edges=p_pca_m$edges,
axes="bottomleft",
edges.col="#E7950F",
edges.width=3)
render_gif(p_pca_m_d,
grand_tour(),
display_xy(half_range=4.2,
edges=p_pca_m$edges,
edges.col="#E7950F",
edges.width=3),
gif_file="gifs/p_pca_model.gif",
frames=500,
width=400,
height=400,
loop=FALSE)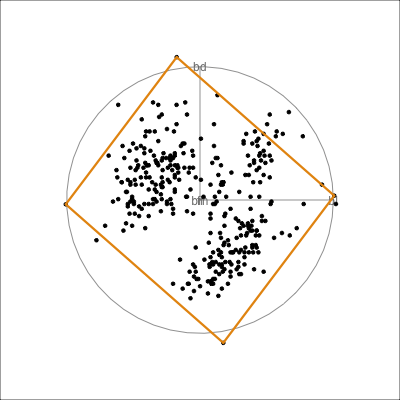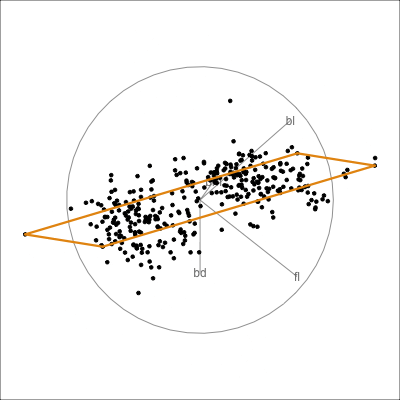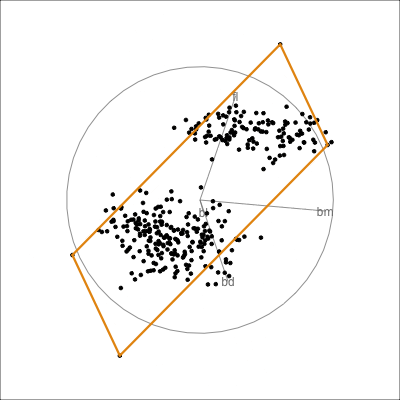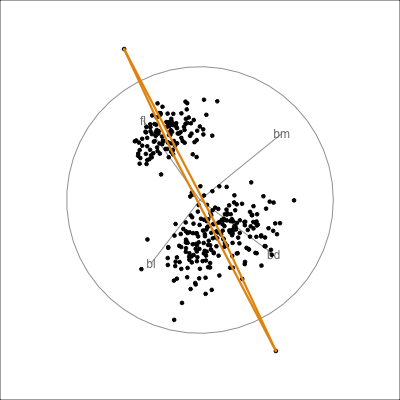
Model in the data space (2/2)
Data in the model space
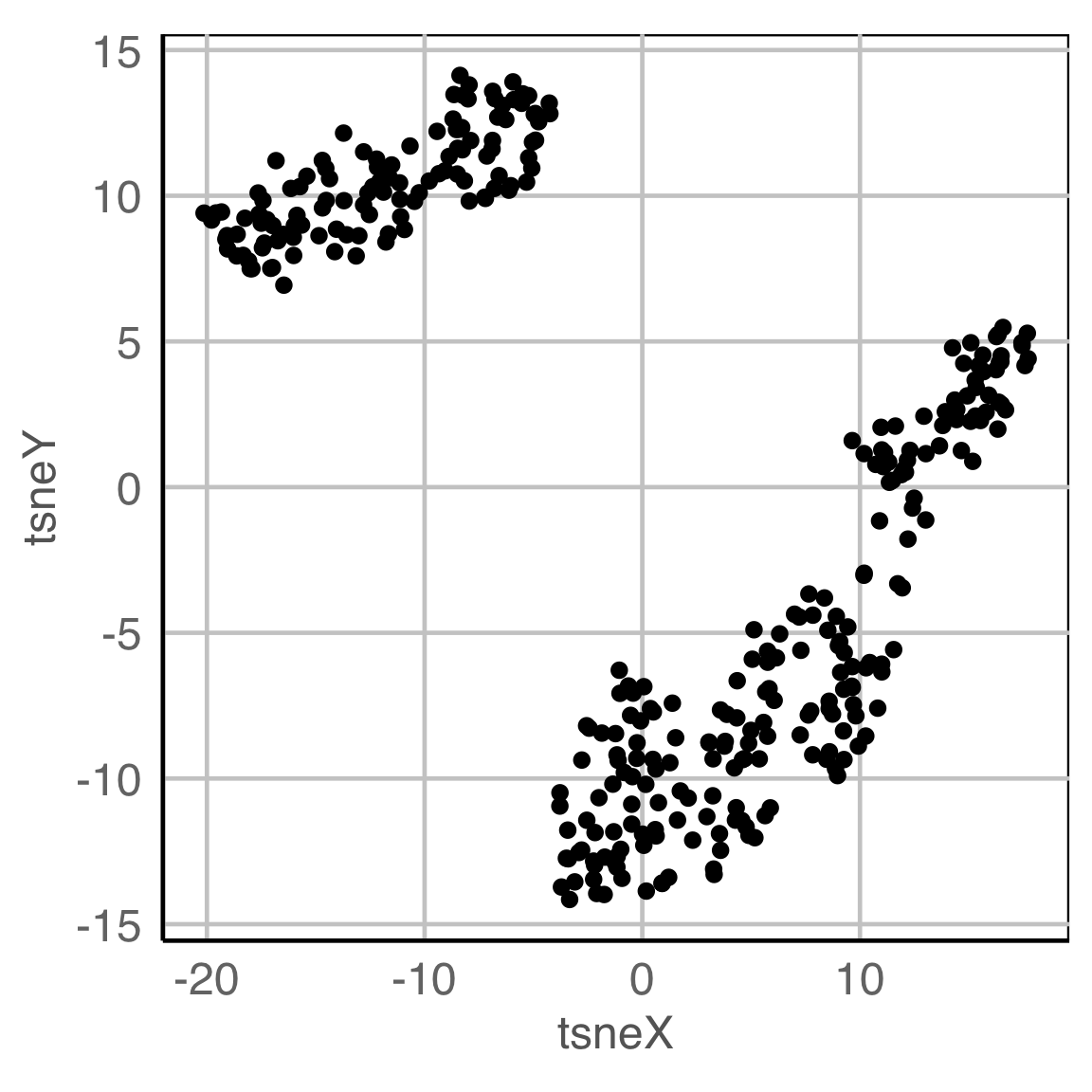
Model in the data space
???
See Jayani Lakshika’s talk, Fri 11am: IPS12
Hiding in high-d (1/2)
Code
library(tidyverse)
library(tourr)
library(GGally)
set.seed(946)
d <- tibble(x1=runif(200, -1, 1),
x2=runif(200, -1, 1),
x3=runif(200, -1, 1))
d <- d %>%
mutate(x4 = x3 + runif(200, -0.1, 0.1))
d <- bind_rows(d, c(x1=0, x2=0, x3=-0.5, x4=0.5))
d_r <- d %>%
mutate(x1 = cos(pi/6)*x1 + sin(pi/6)*x3,
x3 = -sin(pi/6)*x1 + cos(pi/6)*x3,
x2 = cos(pi/6)*x2 + sin(pi/6)*x4,
x4 = -sin(pi/6)*x2 + cos(pi/6)*x4)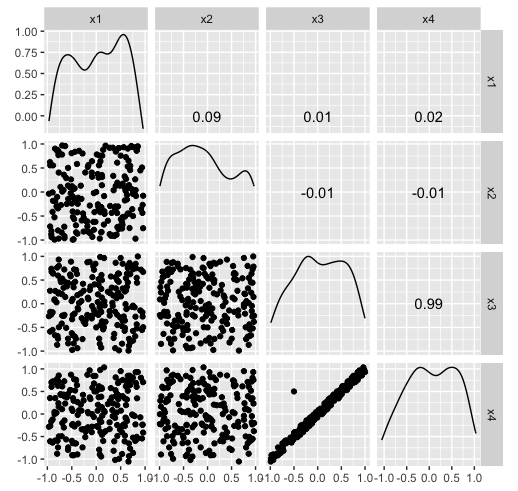
Hiding in high-d (2/2)
Code
library(tidyverse)
library(tourr)
library(GGally)
set.seed(946)
d <- tibble(x1=runif(200, -1, 1),
x2=runif(200, -1, 1),
x3=runif(200, -1, 1))
d <- d %>%
mutate(x4 = x3 + runif(200, -0.1, 0.1))
d <- bind_rows(d, c(x1=0, x2=0, x3=-0.5, x4=0.5))
d_r <- d %>%
mutate(x1 = cos(pi/6)*x1 + sin(pi/6)*x3,
x3 = -sin(pi/6)*x1 + cos(pi/6)*x3,
x2 = cos(pi/6)*x2 + sin(pi/6)*x4,
x4 = -sin(pi/6)*x2 + cos(pi/6)*x4)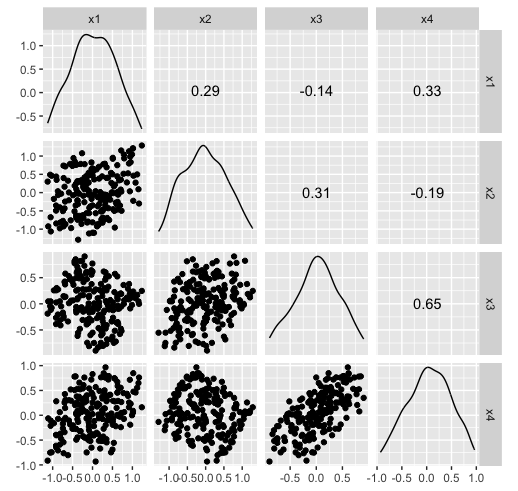

Future work, possible research
The tourr package provides the algorithm to generate the tour paths, and also create new tours, different displays.
- Stopping, pausing, going back
- Zooming in, focus on subsets
- Linking between multiple displays
Elegant interactivity solutions with detourr, liminal, langevitour but need to be developed further.
- Better integration with model objects
- Specialist design for different models
- Integrating other guidance, explainability metrics
High-d vis intellectually challenging, and fun!
Talks at this conference to learn more:
- Fri 11am: IPS12: Visualising high-dimensional and complex data (Paul Harrison, Jayani Lakshika)
- Fri 1:30pm: CPS09: Visualising Complex data & Anomaly Detection (Janith Wanniarachchi - XAI)
Please use these tools at home 😃
References and acknowledgements
- Cook and Laa (2023) Interactively exploring high-dimensional data and models in R
- Slides made in Quarto
- Get a copy of slides at
https://github.com/dicook/IASC-ARS-2023

This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.
Clustering & tours
Model-based - 2D (1/3)
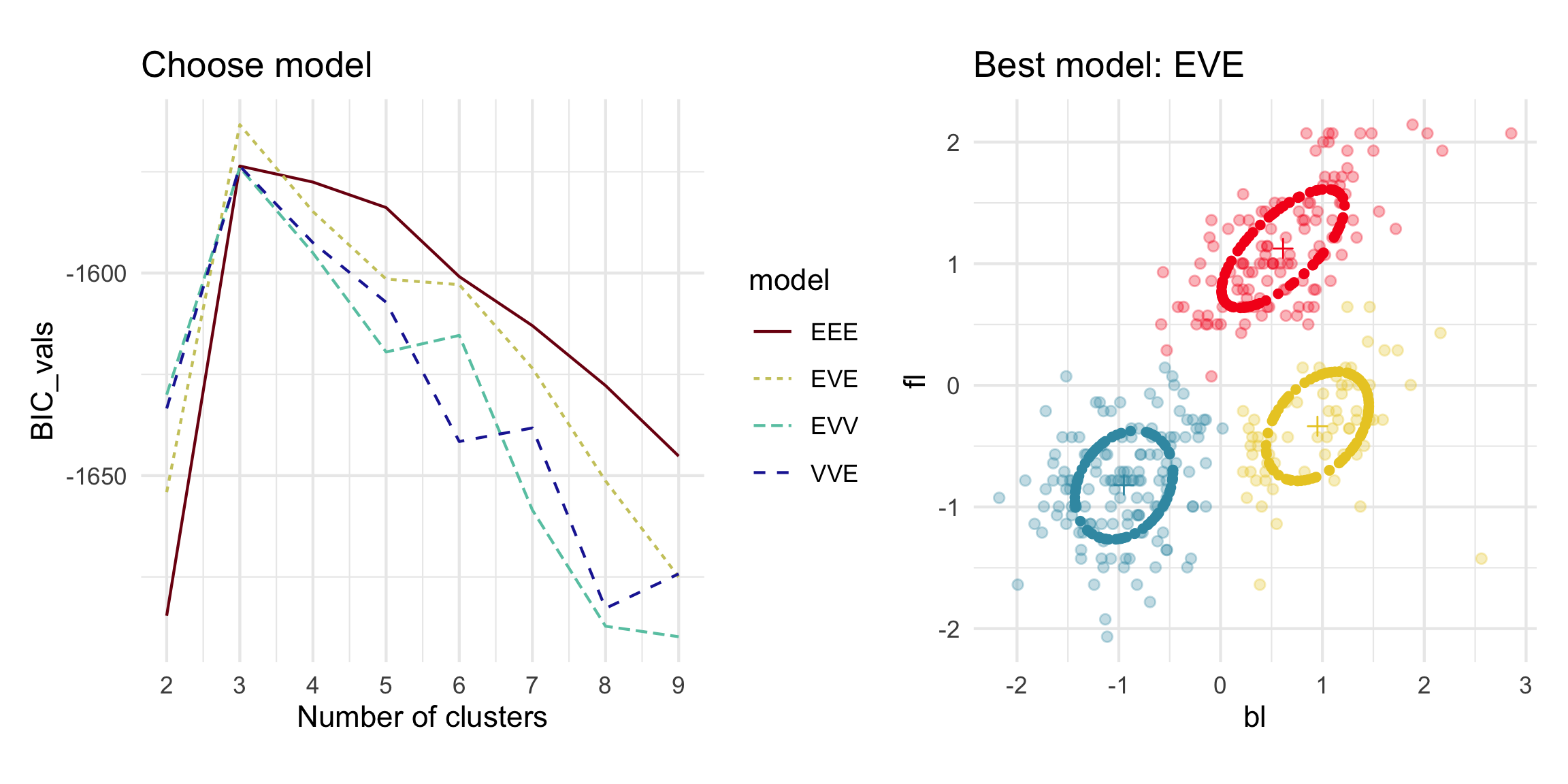
Model-based - 4D (2/3)
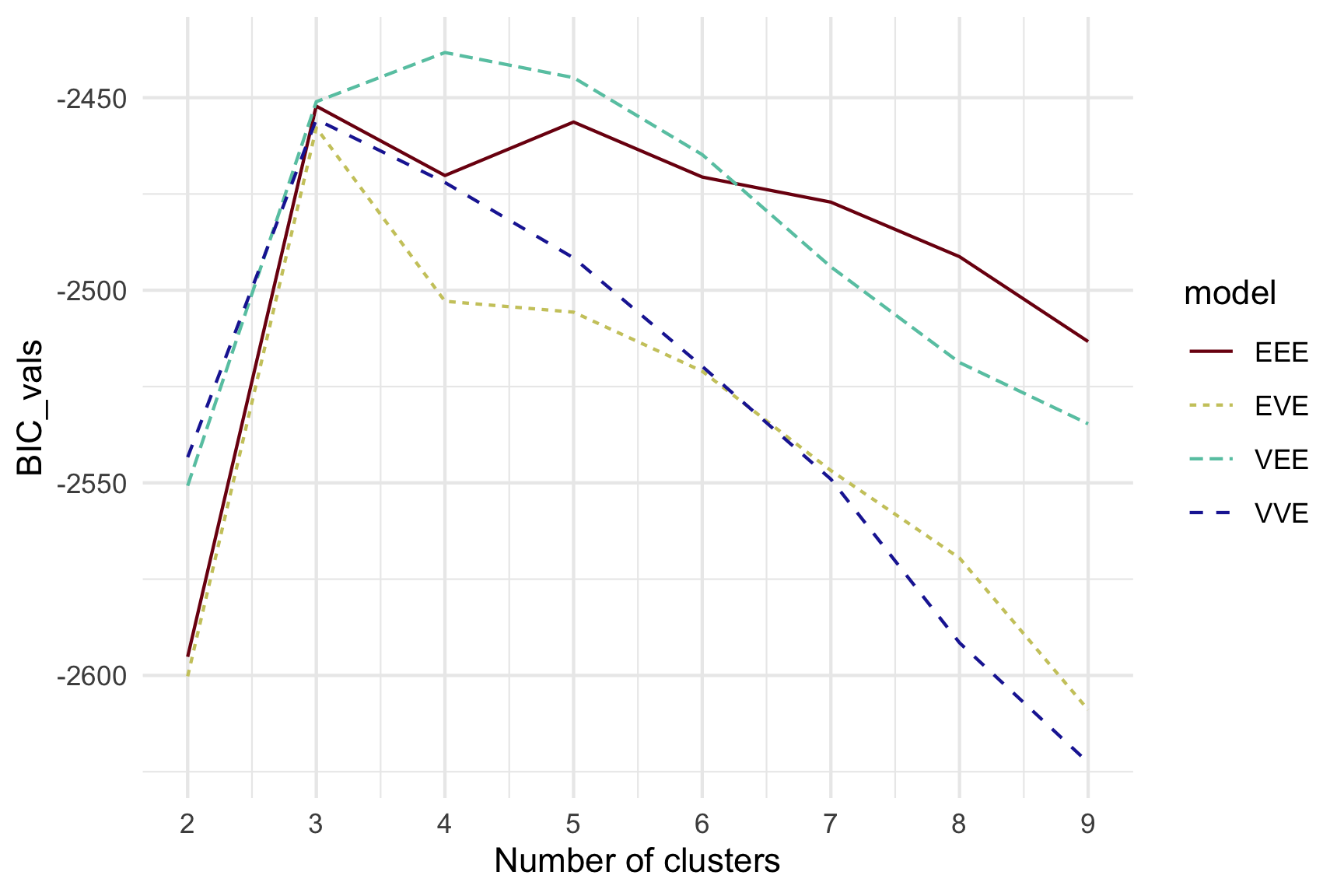
Model-based (3/3) ~~Which fits the data better?
Best model: four-cluster VEE
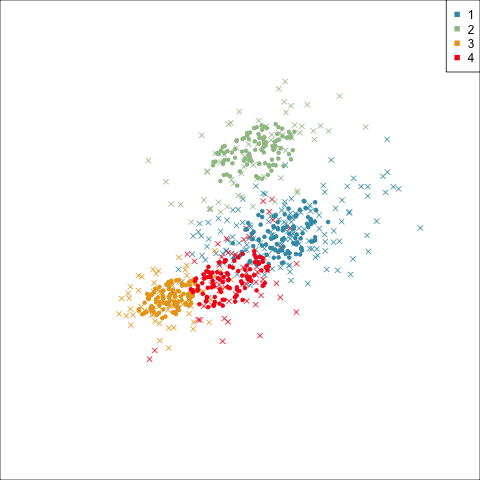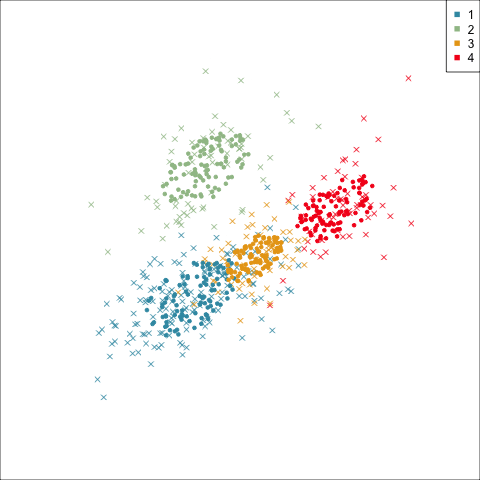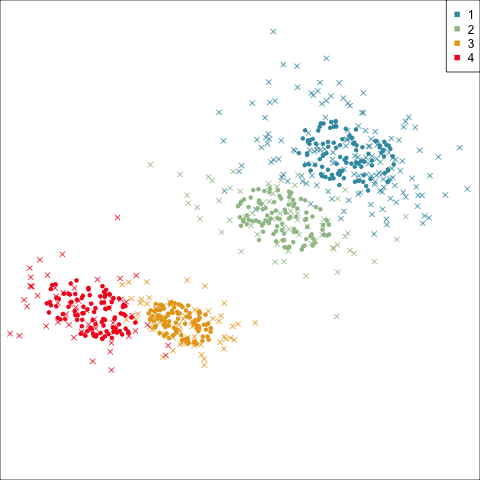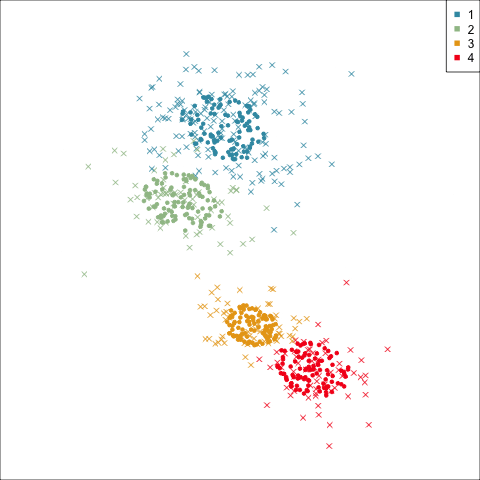
Three-cluster EEE
Summarising clusters
Convex hulls are often used to summarise clusters in 2D. It is possible to view these in high-d, too.