library(flexclust)
randomTour(iris[,1:4], axiscol=2:5)Interactively Visualizing Multivariate Market Segmentation Using the R Package Lionfish
Econometrics and Business Statistics
Monash University
Joint with Ursula Laa and Matthias Medl, BOKU
Fritz’s work
It turns out that Fritz implemented a tour:
Today’s work extends it with better algorithms for choosing projections to show, and interactive graphics where plots are linked in python.

Motivation
“You can get any result you want when clustering data.”
Yes, and no, ideally no
Market segmentation tends to be carving a “blob” of data into chunks, using clustering algorithms. We argue that:
- Clustering follows the shape of the data along mathematical rules
- Algorithms have favourites and quirks, which is replicable and repeatable






Objective
Learn about the shape of the data and how a clustering has carved up the data …
… by using tours - linear projections of high-dimensional data.
Quick quiz
This is how we tend to visualise cluster results.
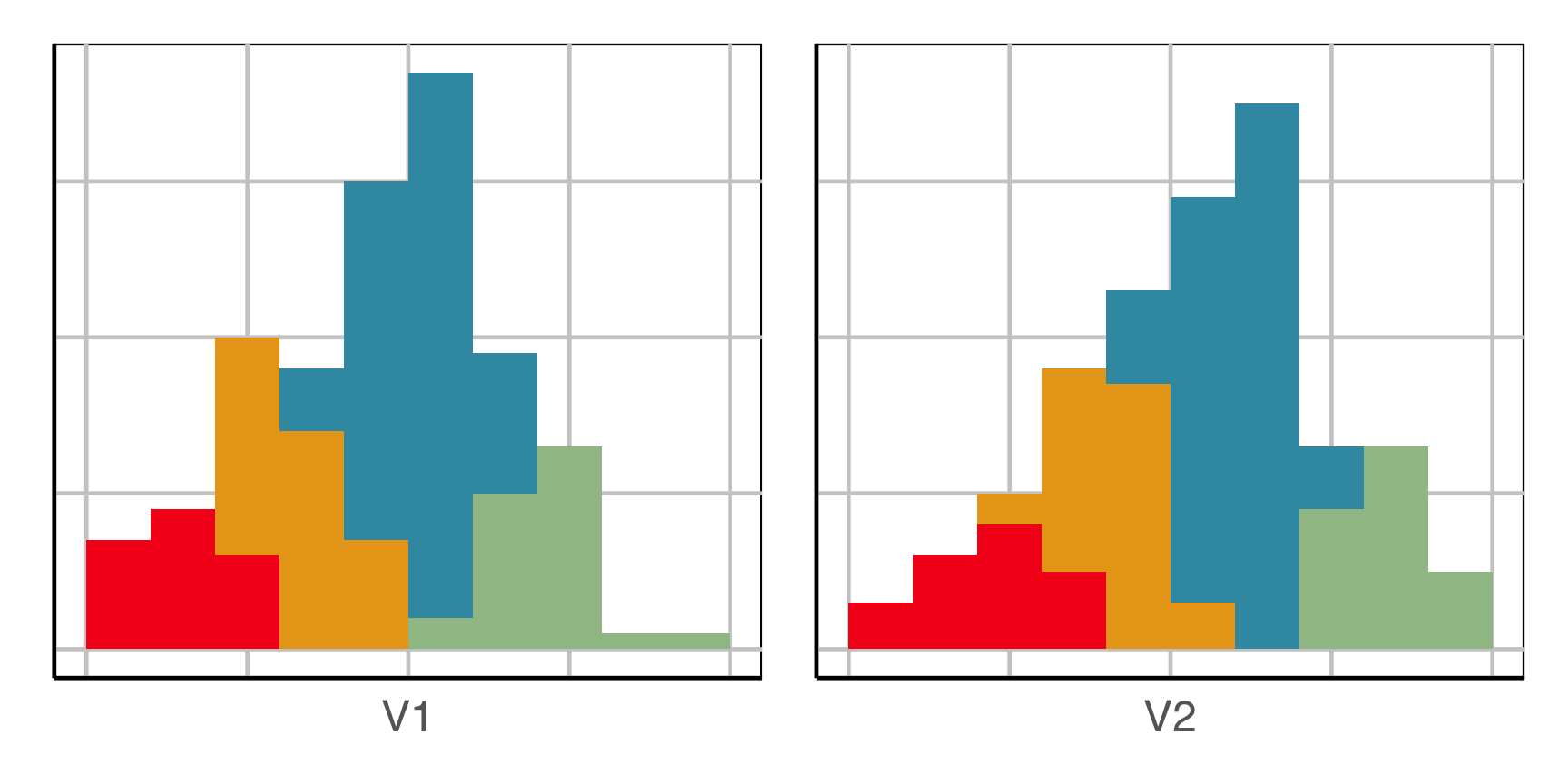
How does this clustering result carve up 2D data? What does the data look like?
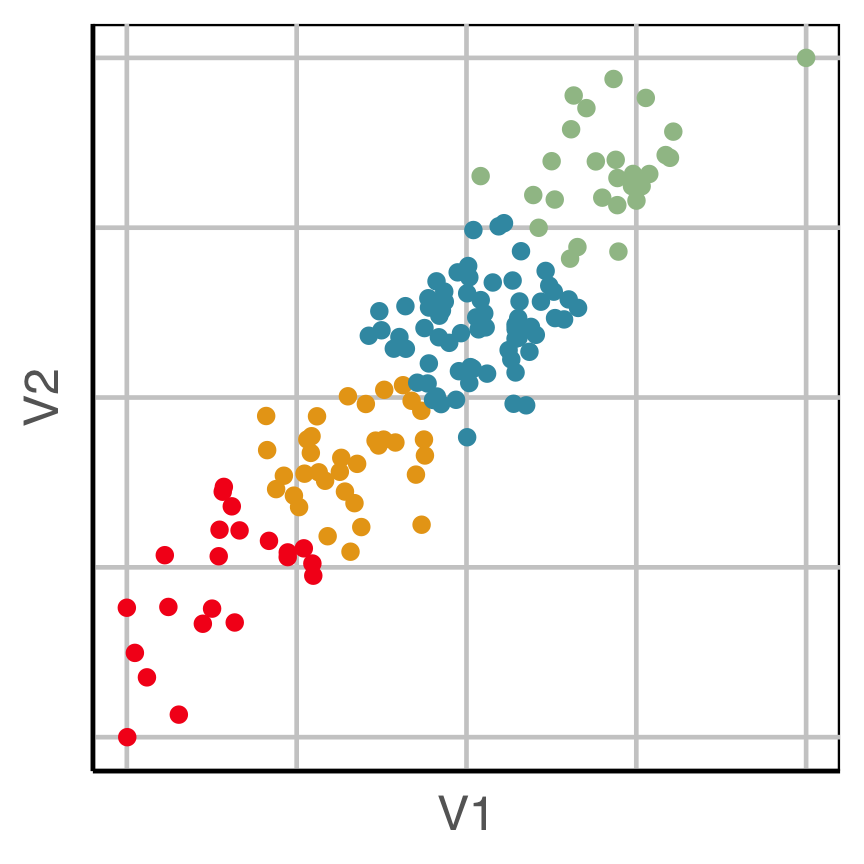
Now we can see the clustering has partitioned the blob.
Try again
This is how we tend to visualise cluster results.
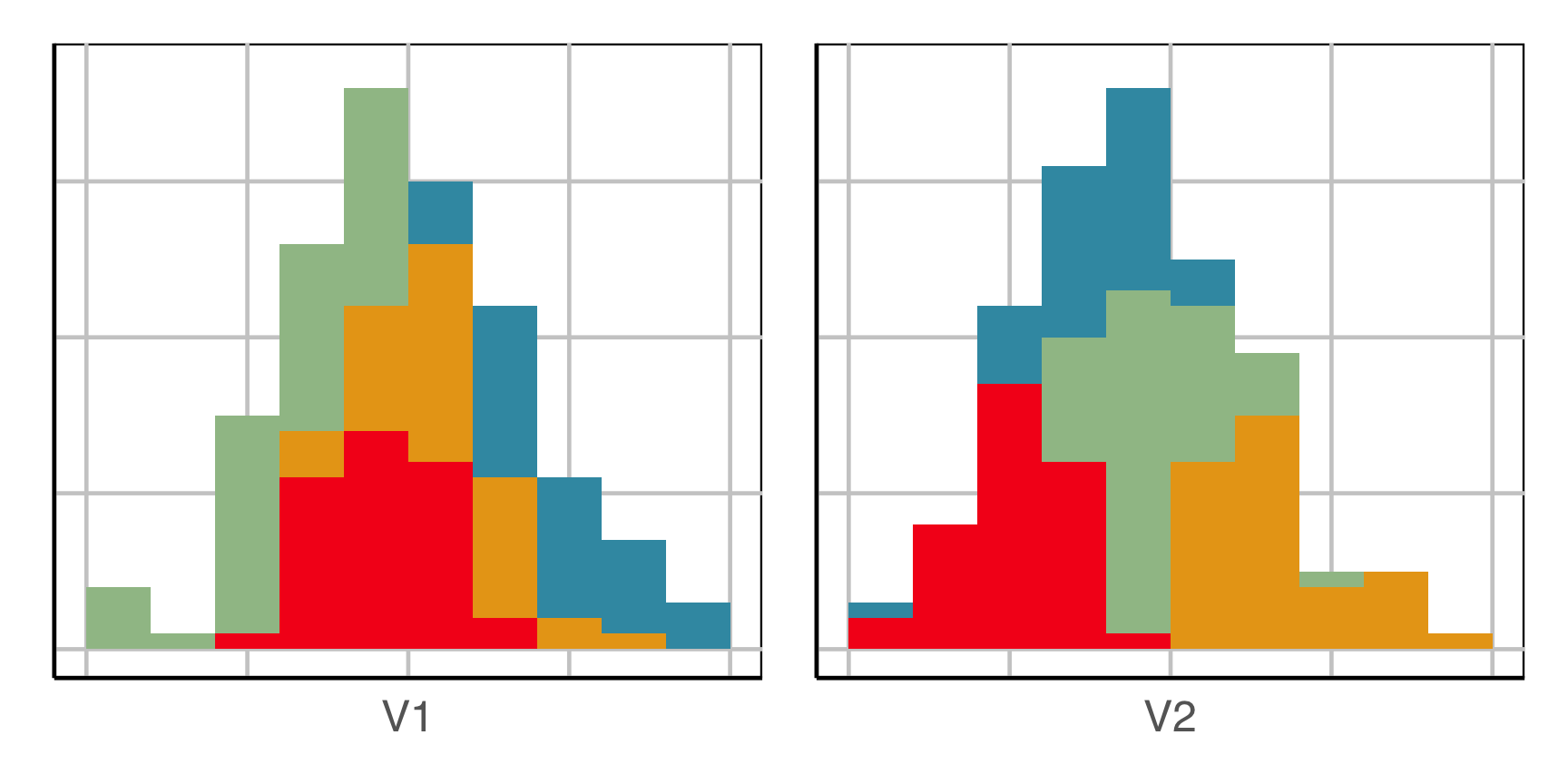
How does this clustering result carve up 2D data? What does the data look like?
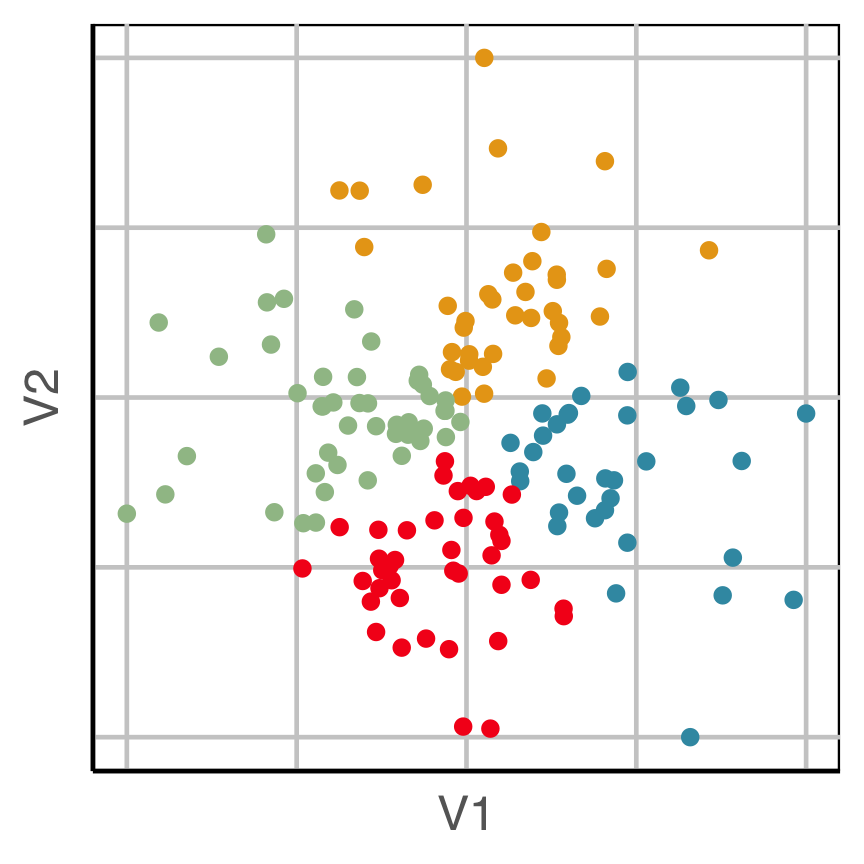
Now we can see the clustering has partitioned the blob.
Searching for the partitions in high dimensions
Example: Risk Taking
- Survey of 563 Australian tourists, see Dolnicar S, Grün B, Leisch F (2018)
- Six different types of risks: recreational, health, career, financial, social and safety
- Rated on a scale from 1 (never) to 5 (very often)
Step 1: understand the shape of the data
Code
# Step 1: get a sense of the data
library(lionfish)
data("risk")
colnames(risk) <- c("Rec", "Hea", "Car", "Fin", "Saf", "Soc")
animate_xy(risk)
set.seed(201)
render_gif(risk,
grand_tour(),
display_xy(col = "#6C26AC"),
start = basis_random(6,2),
gif_file = "gifs/risk_gt.gif",
apf = 1/20,
frames = 400,
width = 400,
height = 400)

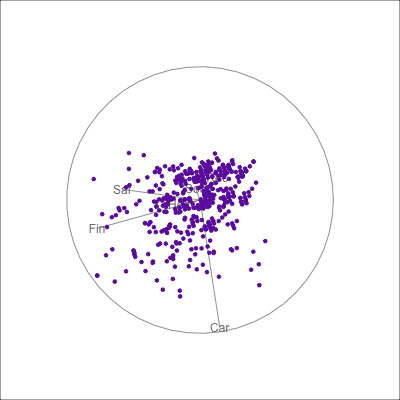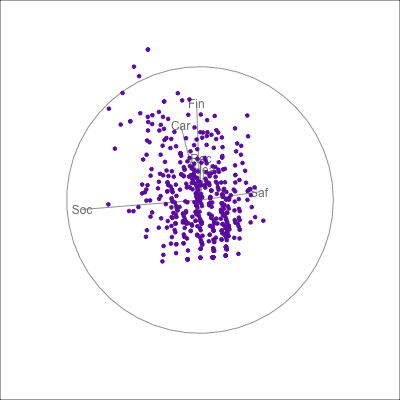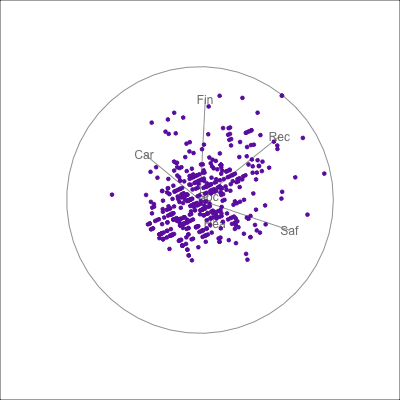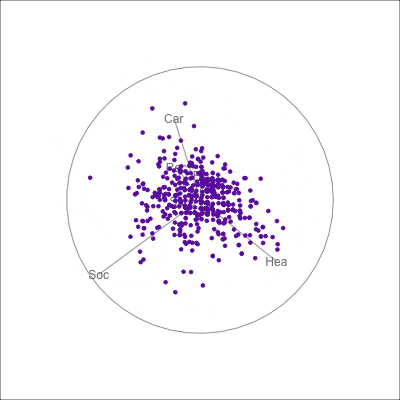
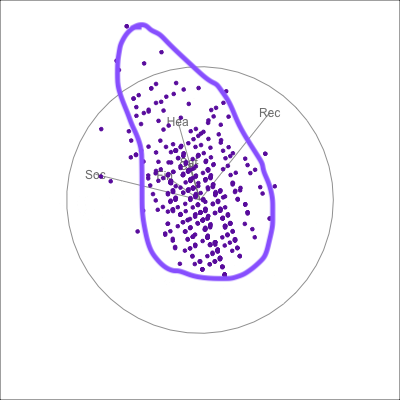 {fig-alt=“A single 2D projection of 6D data shown as a scatterplot of purple points. A purple sketch roughs out the shape, which is like a pear. The variables are mostly contributing to this projection Soc, Rec and Hea.”}
{fig-alt=“A single 2D projection of 6D data shown as a scatterplot of purple points. A purple sketch roughs out the shape, which is like a pear. The variables are mostly contributing to this projection Soc, Rec and Hea.”}
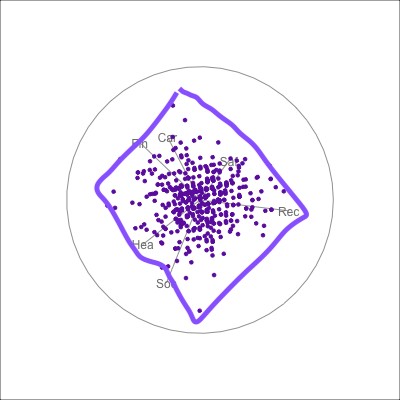 {fig-alt=“A single 2D projection of 6D data shown as a scatterplot of purple points. A purple sketch roughs out the shape, which is like a rhombus. All six variables contribute to this projection in different directions.”}
{fig-alt=“A single 2D projection of 6D data shown as a scatterplot of purple points. A purple sketch roughs out the shape, which is like a rhombus. All six variables contribute to this projection in different directions.”}
Software: lionfish
Rpackage to work with implementations of clustering algorithms, and with thetourrpackage to generate tour pathspythoninterface to useTKinterandmatplotlibfor the GUI and the interactive graphicsmatplotlibenables fast rendering and interactivity for linked brushing and manual tours
Finding the partitions
- Run the clustering
- Run a guided tour with the LDA index to find projection that best separate clusters
- Manual tour to refine view of partitions
Code
# Initialise python environment
init_env()
library(tibble)
library(dplyr)
risk_d <- apply(risk, 2, function(x) (x-mean(x))/sd(x))
# Two clusters
nc <- 3
set.seed(1145)
r_km <- kmeans(risk_d, centers=nc,
iter.max = 500, nstart = 5)
r_km_d <- risk_d |>
as_tibble() |>
mutate(cl = factor(r_km$cluster)) |>
bind_cols(model.matrix(~ as.factor(r_km$cluster) - 1))
colnames(r_km_d)[(ncol(r_km_d)-nc+1):ncol(r_km_d)] <- paste0("cluster", 1:nc)
r_km_d <- r_km_d |>
mutate_at(vars(contains("cluster")), function(x) x+1)
clusters <- r_km_d$cl
set.seed(110)
guided_tour_history <- save_history(risk_d,
tour_path = guided_tour(lda_pp(clusters)))
half_range <- max(sqrt(rowSums(risk_d^2)))
feature_names <- colnames(risk_d)
cluster_names <- LETTERS[1:nc]
clusters <- as.numeric(as.character(clusters))
obj1 <- list(type="2d_tour", obj=guided_tour_history)
risk_d <- data.matrix(risk_d)
interactive_tour(data=risk_d,
plot_objects=list(obj1),
feature_names=feature_names,
half_range=half_range,
n_plot_cols=2,
preselection=clusters,
preselection_names=cluster_names,
n_subsets=nc,
display_size=6)-means slices along main spread, and then middle
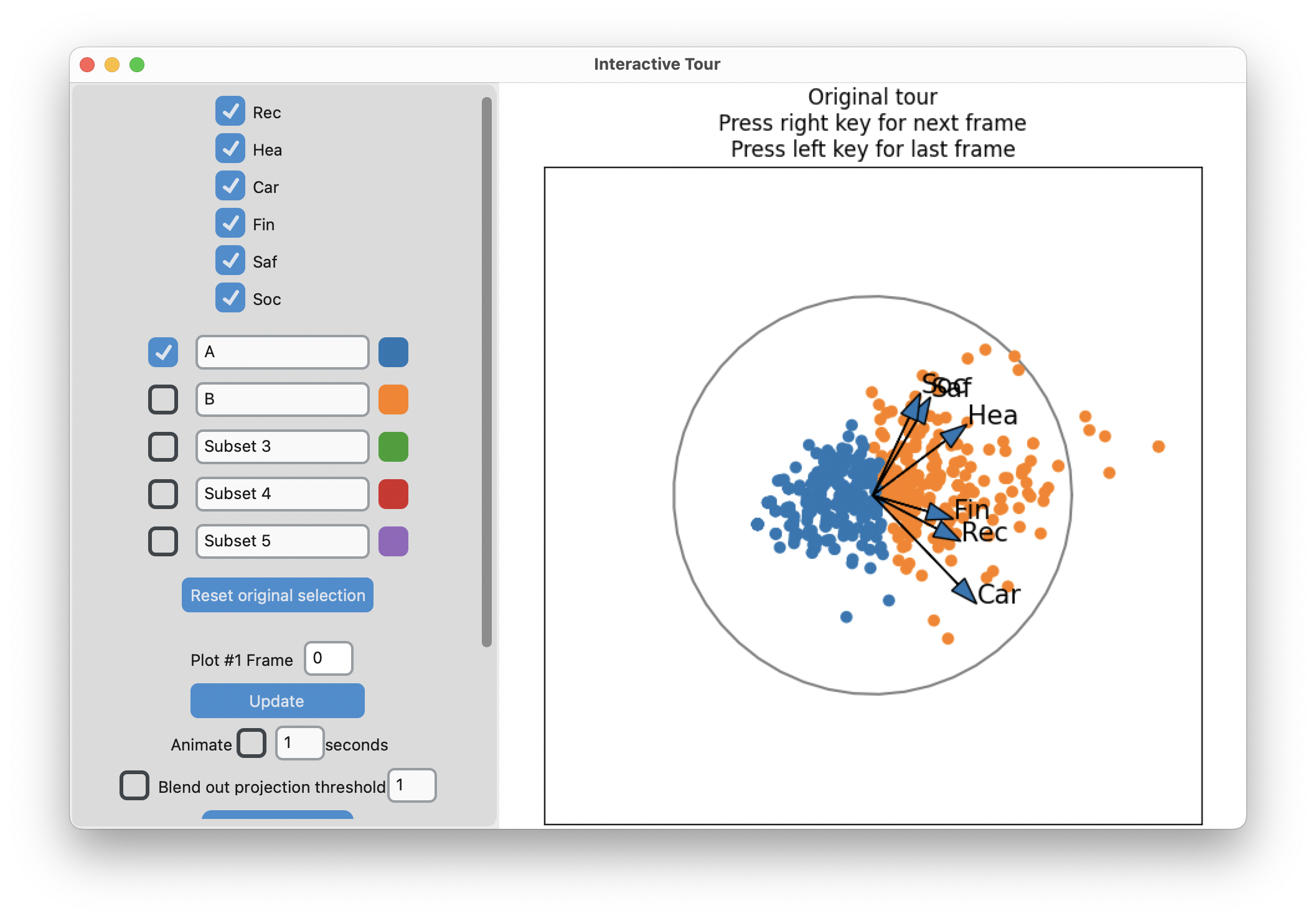
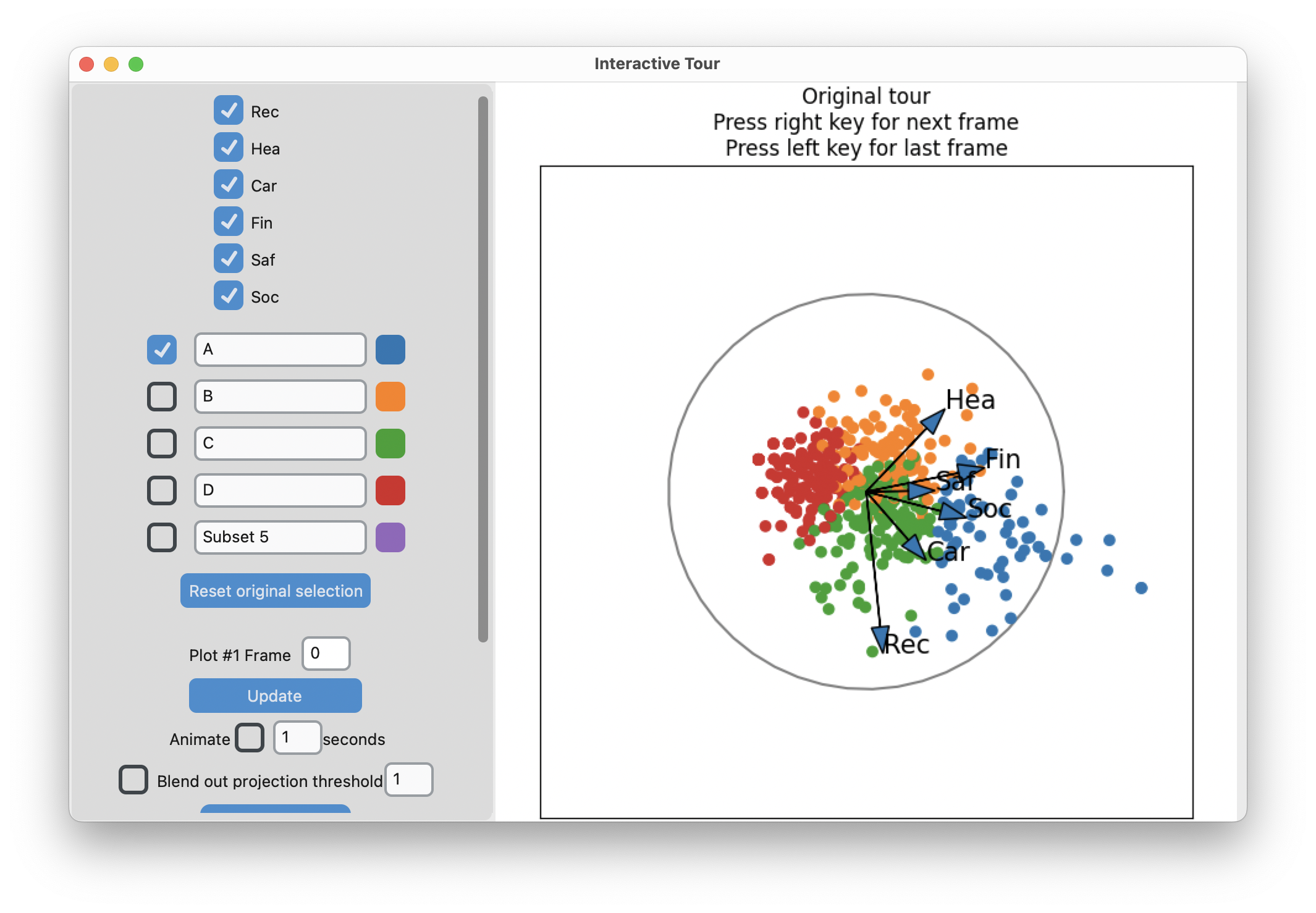
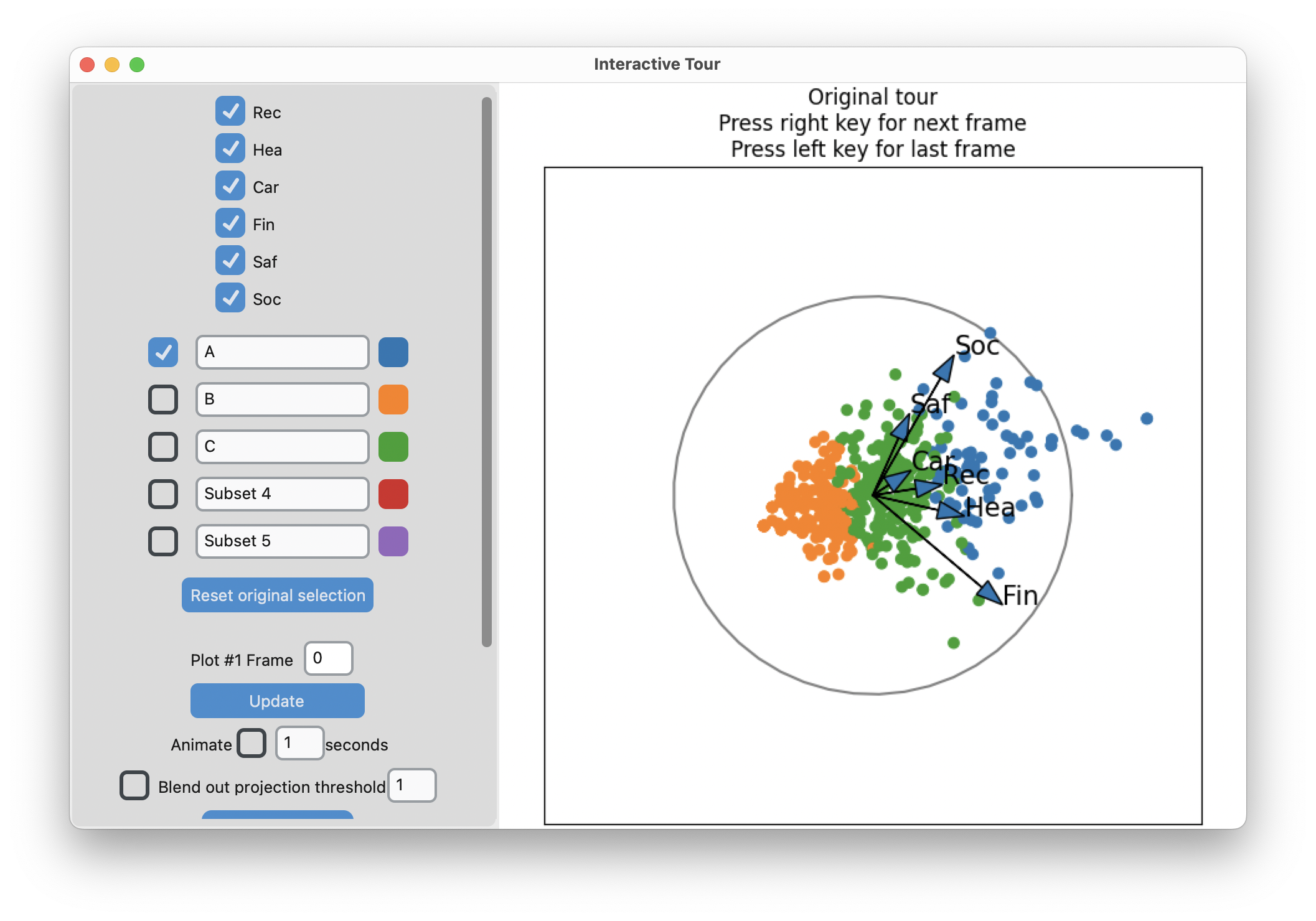
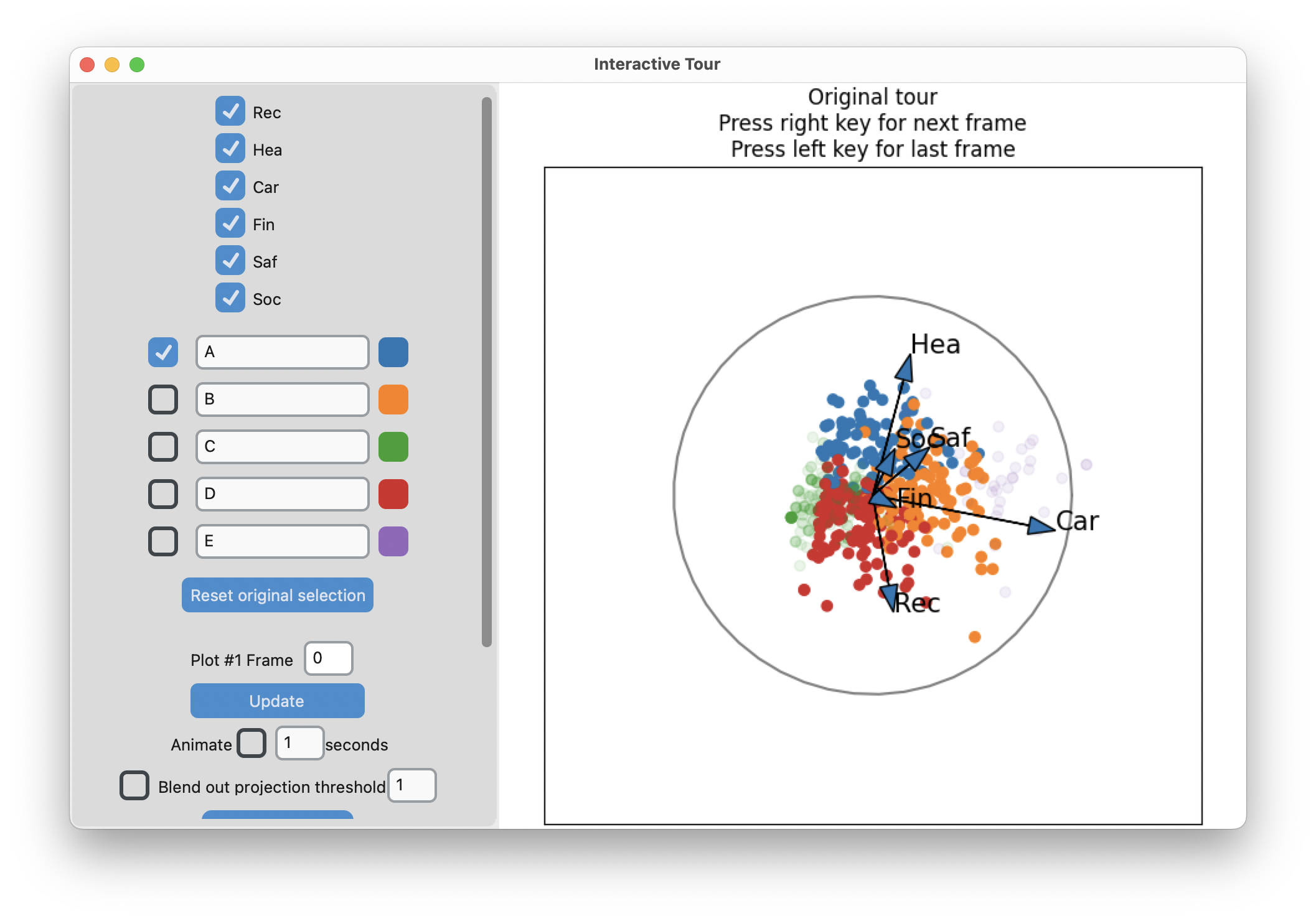
Search for meaning
Code
library(tibble)
library(dplyr)
risk_d <- apply(risk, 2, function(x) (x-mean(x))/sd(x))
# Two clusters
nc <- 3
set.seed(1145)
r_km <- kmeans(risk_d, centers=nc,
iter.max = 500, nstart = 5)
r_km_d <- risk |>
as_tibble() |>
mutate(cl = factor(r_km$cluster))
r_km_d |>
pivot_longer(Rec:Soc, names_to = "var", values_to = "val") |>
ggplot(aes(x=val, fill=cl)) +
geom_bar() +
facet_wrap(~var, ncol=3, scales="free_y") +
scale_fill_manual(values = c("#377EB8", "#FF7F00", "#4DAF4A")) +
xlab("") + ylab("") +
theme_minimal() +
theme(legend.position = "none",
axis.text = element_blank())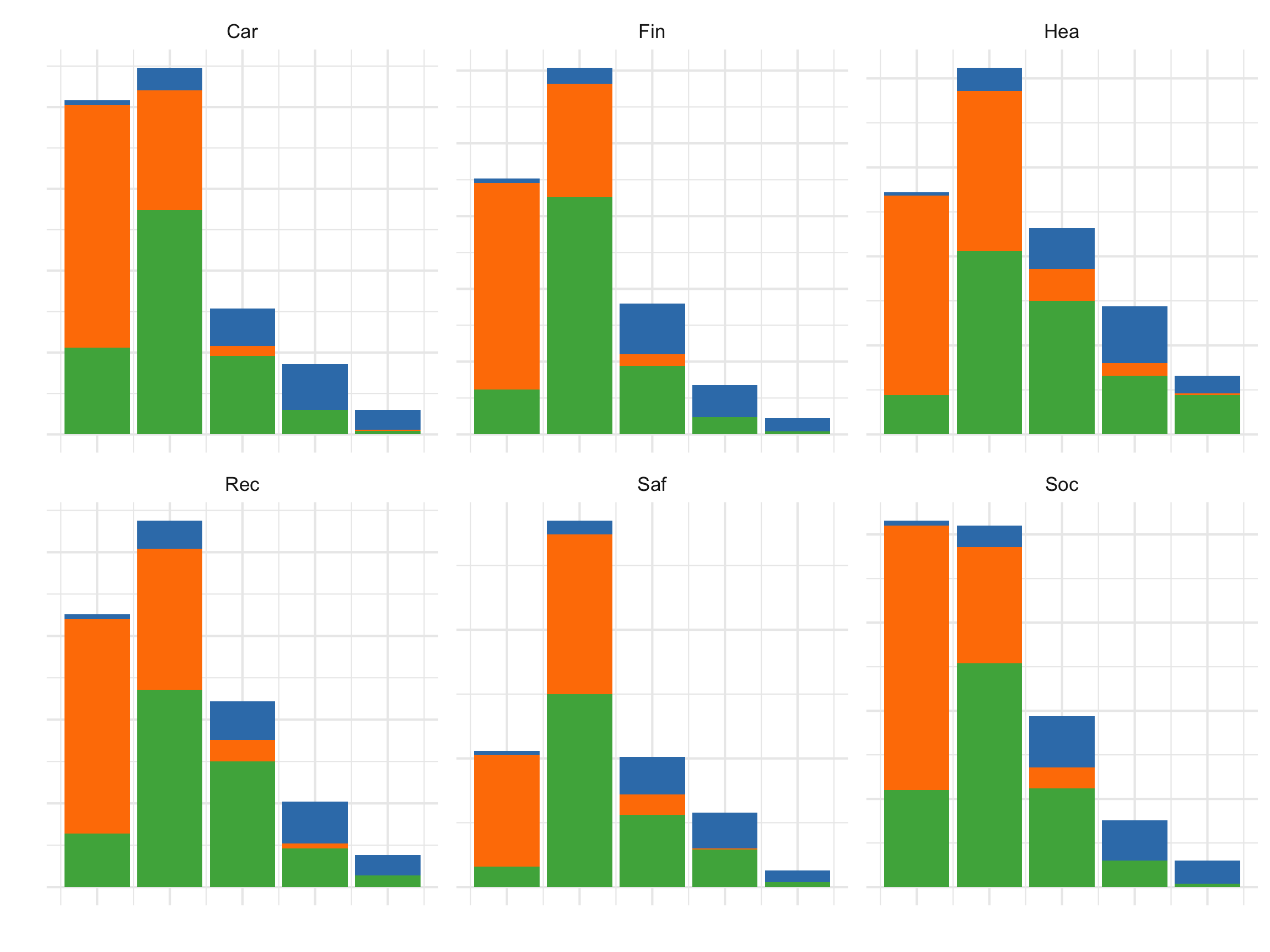
All the activities contribute to the segmentation into three clusters.
Summary
You can find lionfish at https://mmedl94.github.io/lionfish/.
- Link multiple displays
- Interactively select points and clusters
- Visualize the partitions with various tour types
Clustering is a geometric operation and using the tour you should be able to see how the observations have been grouped, always.
Final teaser: clustering algorithms don’t see gaps, it sees pairwise distances. We see gaps, and can be shocked when the algorithm grouped across it.
References and acknowledgements
- Medl, Cook, Laa (2025) Demonstrating the Capabilities of the Lionfish Software for Interactive Visualization of Market Segmentation Partitions
- Cook and Laa (2025) Interactively exploring high-dimensional data and models in R
- Wickham et al (2015) Visualizing statistical models: Removing the blindfold
- Flatland: A Romance of Many Dimensions (1884) Edwin Abbott
Slides made in Quarto, with code included.

This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.
Symposium in Memory of Fritz Leisch - https://github.com/dicook/Leisch_Symp_2025